Journal of Plant Biology & Soil Health
Download PDF
Research Article
Isolation, Characterization and Genetic Studies on Isolates of Phosphate Solubilizing Bacteria in Egyptian Calcareous Soils
Abdelaziz S1*, Hemeda NF2, Belal EE3 and Serag AM4
- 1Department of Agricultural Microbiology, Fayoum University, Egypt
- 2Department of Genetics, Fayoum University, Egypt
- 3Department of Soils and Water, Fayoum University, Egypt
- 4Department of Genetics and Genetic Engineering, Benha University, Egypt
*Abdelaziz S, Faculty of Agriculture, Department of Agricultural Microbiology, Fayoum University, Egypt, E-mail: sa171@fayoum.edu.eg
Citation: Abdelaziz S, Hemeda NF, Belal EE, Serag AM. Isolation, Characterization and Genetic Studies on Isolates of Phosphate Solubilizing Bacteria in Egyptian Calcareous Soils. J Plant Biol Soil Health. 2019;6(1): 10.
Copyright: © 2019 Abdelaziz S, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Journal of Plant Biology & Soil Health | ISSN: 2331-8996 | Volume: 6, Issue: 1
Submission: 10 January, 2019 | Accepted: 13 Februay, 2019 | Published: 15 Februay, 2019
Abstract
Phosphorous (P) is an essential nutrient element and plays an important role in plant growth and development, it mostly presented in form unavailable for plants. Phosphate Solubilizing Bacteria (PSB) can be successfully used for solubilizing such forms rendering them available for plants. Thirty-two PSB strains were isolated on a Pikovskaya (PKV) agar medium containing Tricalcium Phosphate (TCP) and examined for plant growth promoting effects. A high portion of isolates (68.8%) produced Indole Acetic Acid (IAA) in contents ranging from 5 to 15 μgmL-1 and 12.5% produced salicylic acid (SA) in contents < 100 μgmL-1 while 50.0% fixed gaseous N2 nitrogen in medium deprived completely of nitrogen. A portion of 28.1% produced cellulose enzyme and 15.6% produced chitinase enzyme. In vitro tests showed that isolates were capable in controlling some fungus plant pathogens and isolates were resistance to some adverse conditions involving pH, temperature and salinity. Use of 16s rDNA analysis and other procedures showed that the most 3 effective isolates were Bacillus megaterium-MH142578, Acinetobacter lwoffii-MH142579 and Acinetobacter lwoffii-MH142580. The results of cluster analysis (Similarity index) showed that were high and low similar values between the bacterial genera under studies.
Keywords
Phosphate solubilizing bacteria; Phosphate uptake plant growth promoting traits; 16s rDNA; Phylogenetic and calcareous soils
Introduction
Phosphate (P) is one of the most important elements for plant growth. Although it is abundant in soils in both organic and inorganic forms, its availability is limited to plants as it presents mostly in insoluble forms [1]. Chemical fertilizers are added to the soils, plants can only utilize few amounts of phosphoric fertilizers, which are quickly converted into insoluble forms. Consequently, chemical fertilizers are frequently applied during crop planting, but its regular use is costly and produces undesirable environmental impacts, such as soil and water contamination. Therefore, P is often regarded a limiting nutrient in agricultural soils [2,3]. Soil microorganisms play a great role in availability of phosphate to plants [4].
Biofertilizers are considered to be the alternate source to meet the nutrient requirement of crops and to bridge the future gaps. Soil microorganisms which can solubilize phosphorus, fix atmospheric nitrogen or stimulate plant growth through synthesis of growth promoting substances could be used as safe alternative for the overuse of harmful agrochemicals [5-7]. Soil microorganisms play a great role in availability of phosphate to plants [8]. Microorganisms have the capacity to solubilize Phosphate, These include bacteria, fungi, actinomycetes and algae, and they are collectively called as Phosphate Solubilizing Microorganisms. Bacteria are more successful in P solubilization than others. The Plant Growth Promoting Rhizobacteria (PGPR) include Phosphate solubilizing bacteria [9,10].
Consequently, the present work aims to study the prevalence of the phosphate solubilizing bacteria in various soils of Fayoum governorate, isolation of some individuals and studying some of their characters and activities (N2-fixation, siderophores, Salicylic Acid (SA) and Indole Acetic Acid (IAA) production. The capability of these isolates in controlling some plant pathogenic fungi (in vitro) was also determined. The resistibility of isolates to some adverse conditions prevailing in our conditions (pH, temp. and salt content) was also included. The isolates were identified by using 16s rDNA analysis after DNA bacteria isolation and the most 3 effective isolates Bacillus megaterium-MH142578, Acinetobacter lwoffii-MH142579 and Acinetobacter lwoffii-MH142580 were involved on proceeding study. Generally strains of PSB have plant growth-promoting activities and antagonistic potential against phytopathogenic fungi that could be used as alternate source for the overuse of harmful agrochemicals.
Materials and Methods
Collection and characterization of soils samples
Soil samples were collected from two sites distinguished by calcareous soils conditions, site 1 in Youssef El-Sediek and site 2 in Domo in Fayoum Governorate area; surface soil sample (0-30 cm) was collected from the two sites. Collected soil samples were airdried, gently crushed and passed through a 2 mm sieve and stored in plastic bottles. Some physical and chemical properties of the soil were analyzed. Particle size distribution of the collected soil samples were carried out by the international pipette method [11]. Soil texture class was obtained. The pH values of soil samples were measured in the saturated soil paste using Bekman pH meter also Electrical conductivity values were determined in the saturated soil-water paste extract as dS/m, using CM25 conductivity meter [12]. Total calcium carbonates were determined using Collinś Calcimeter [13]. Soil organic matter contents were determined using the wet combustion method according to Walkly and Black ś method [14]. Total nitrogen was determined using micro-kjeldahl method [15]. Available potassium was extracted using 1.0 N ammonium-acetate solution at pH 7.0 [12]. Available P was extracted with sodium bicarbonate 0.5 N (NaHCO3) solution at pH 8.5 [16]. Available micronutrients (Fe, Mn, Zn and Cu) were extracted by DTPA solution [17]. Then measured with Inductively Coupled Plasma (ICP) atomic emission spectroscopy.
Isolation of phosphate solubilizing bacteria
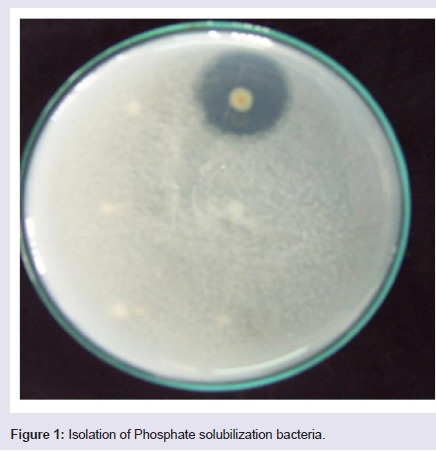
Ten grams of each soil sample was suspended in 90 ml sterile saline solution and was shaken for 1 h. The serially diluted soil samples were placed on Petri dishes containing Pikovskaya’s (PKV) agar medium by pour plate technique and incubated at 28 ± 2 °C for 48-96 h [18]. The bacterial colonies showing clear zone around them were considered as Phosphate Solubilizing Bacteria (PSB) as shown in [19] (Figure 1). Pure culture of the isolates was made by repeated sub culturing for 2-3 times on fresh PKV plate and was maintained on PKV slants at refrigerator temperature.
Assay of solubilization index (SI)
To determine the Solubilization Index (SI), the sterilized Pikovskaya’s (PKV) agar medium was poured into sterilized Petri plates, containing insoluble Ca3(PO4)2 at 5 g/L-1. After solidification of the media, the plates were inoculated with the isolated bacteria, incubated at 28 ± 2 °C for two weeks and assayed visually. The solubilization index was determined by measuring the halo (clear zone) diameter and the colony diameter, using the following formula [20]. All assays were replicated three times.
Formula Morphological, physiological and biochemical characters of isolates
Different morphological characters of isolates obtained were observed on NB agar media incubated for 48 hours at 30 °C. Cell shape was examined microscopically using gram stain in addition to motility. Catalase and oxidase activities were measured also.
Characterization of plant growth prompting traits
The capability to produce Plant Growth Promoting substances (PGPs) such as Indoleacetic Acid (IAA) production was measured by the colorimetric method [21], Siderophores production by different isolates was detected using the method described by Schwyn with several modification by Pallai [22, 23]. Salicylic acid (SA), isolates were grown at 28 °C for 48h on a rotary shaker at 200 rpm in flasks containing 25 ml of the standard succinate medium, the absorbance of the purple iron-SA complex developed in the aqueous phase was measured at 527 nm, using SA dissolved in the growth medium and treated as described above as a control [24, 25]. Cellulose production was visualized by flooding the cellulose decomposition medium plates with 0.1% (w/v) Congo red for 15 to 30 min followed by bleaching the plates with 1 M NaCl, according to the method of [26]. Chitinase production was assessed qualitatively by a microbiological method based on growth of isolates in chitinase medium amended with colloidal chitin [27]. The isolates were screened in vitro for N2 fixation by growing them on plates of N-free agar medium for 48 h at 28 °C. The isolates that grow after being sequentially transferred 10 times to the same medium were considered presumptive positive for N2 fixation [28-30].
Antifungal activity assay
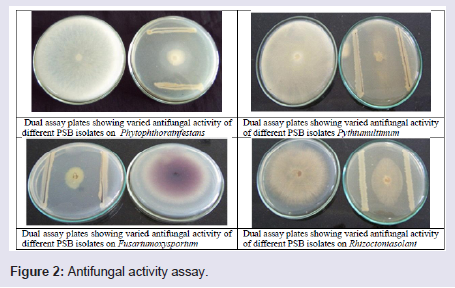
Phosphate Solubilizing Bacteria (PSB) isolates were screened in vitro for antagonism towards four soil borne pathogenic fungi: Pythium ultimum, Rhizoctonia solani, Fusarium oxysporum and Phytophthora infestans in dual culture test [31]. Five μl of an exponentially growing bacterial culture was streaked along two opposite sides of surface-dried Potato Dextrose Agar (PDA) plates. Plates subsequently incubated at 28 °C for 24 h. Following bacterial growth, mycelial agar disc of 5-mm-diameter from a 7-day-old culture from the target fungi grown on PDA plate was placed in the center of the plate between the two parallel streaks of the test bacteria. Plates inoculated with target fungi alone served as control plates, and two replicate plates were used for each bacterial isolate. Plates were then incubated at 25 °C for 7 days. In vitro antagonistic activity was assessed by relating mycelial diameter on plates inoculated with bacteria to mycelial diameter on control plates and computing percentage inhibition as shown in (Figure 2).
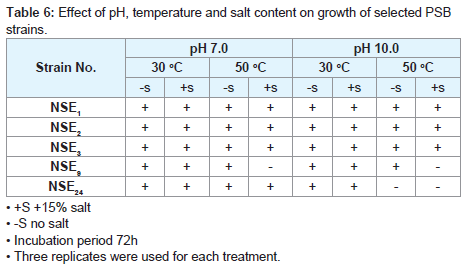
Effect of pH, temperature and salt content on growth of selected strains
The PSB isolates were examined against some adverse environmental conditions to determine their capability to live, proliferate sustain life and perform their activities. They were tested against temperature, increasing pH and salt tolerant characters which prevail in our conditions. They were inoculated at fixed counts on 100 ml complete nutrient broth medium in 250 ml flasks which incubated for 24, 48 and 72 hours at elevated pH, temperature and salt content. Each character was studied separately and in combination with one or two of the other characters. Each treatment was replicated threetimes.
Genotypic characterization by 16s-rDNA
Bacterial DNA extraction and PCR amplification
The nucleic acid was extracted using protocol of beat beading with zirconia silica beads combined with ABT genomic DNA mini extraction kit spin column (Applied biotechnology, Egypt) according to the manufacture’s instruction. A 16s rDNA reference was considered as full-length if it covered the position of the primer (27 F 5’-AGAGTTTGATCCTGGCTCAG-3’ and 1492 R 5--GGTTACCTTGTTACGACTT-3’), used in a previous study [32]. The PCR was performed in the thermal cycle 2720 (Applied Biosystem, USA) in a total volume of 25 μl containing 1 μl of each primer, 200 μM from the four ribonucleotide triphosphates (dNTPs), 5 μl of 10X PCR buffer, 1 μl of 25 m mol Mgcl2, 1 μl of template DNA, 1 μl of Taq DNA polymerase and 14.5 μl of water nuclease-free.
Amplification of DNA was carried out under the following conditions: denaturation at 94 °C for 5 min followed by 30 cycles of 94 °C for 30 s, 52 °C for 30 s, 72 °C for 1.5 min and final extension at 72 °C for 10 min. PCR products of bacterial isolates were checked by electrophoresis with 1.8% agarose gel and stained by ethidium bromide and then visualized and photographed under UV transilluminator. The PCR have specific bands at size around 1500 bp. The amplified PCR product was purified using Montage PCR Clean up Kit (Millipore), following manufacture instructions to remove unincorporated PCR primers.
Sequencing of 16s rDNA and Phylogenetic analysis
The purified PCR products were subjected to sequencing through Solvent Sequencing Service located in Korea. The sequences of 16s rDNA of the three bacteria isolates were submitted to GenBank.
The data of the nucleotide sequence of the ITS regions of rDNA obtained from the three isolates were compared with 16s rDNA collected from Bacillus megaterium and Acinetobacter lwoffii sequences available in GenBank database (http://www.ncbi.nlm.nih.gov/ Blast). NCBI, Bethesda, MD, (USA) [33,34]. The 16s rDNA sequences of all bacterial isolates were aligned with reference sequences showing sequence homology from the NCBI database using Clustal omega multiple sequence alignment. Phylogenetic dendogram was constructed by the clustal omega multiple sequence alignment programs (successor of Clustal W [35]. Phylogenetic trees were constructed by the Un-weighted Pair Group Method with Arithmetic Mean (UPGMA).
Results
Chemical and physical properties of the studied soils
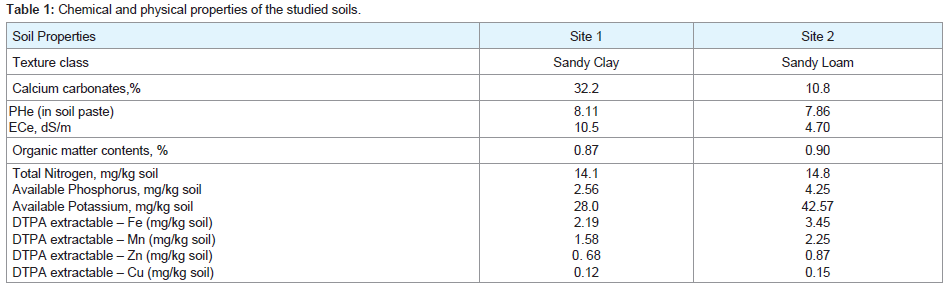
(Table 1) illustrates the available characters of the two samples collected from various locations distinguished by calcareous soils conditions. The soils Chemical and physical properties studied as calcium carbonates %, pHe (in soil paste) ECe dS/m, organic matter contents%, Total Nitrogen mg/kg soil, available Phosphorus mg/kg soil, available Potassium mg/kg soil, available micronutrients such as (Fe, Mn, Zn, and Cu mg/kg soil). Soil texture classes of all samples were diverse between sand clay and sandy loam.
Isolation of phosphate solubilizing bacteria (PSB) and determination of their characters
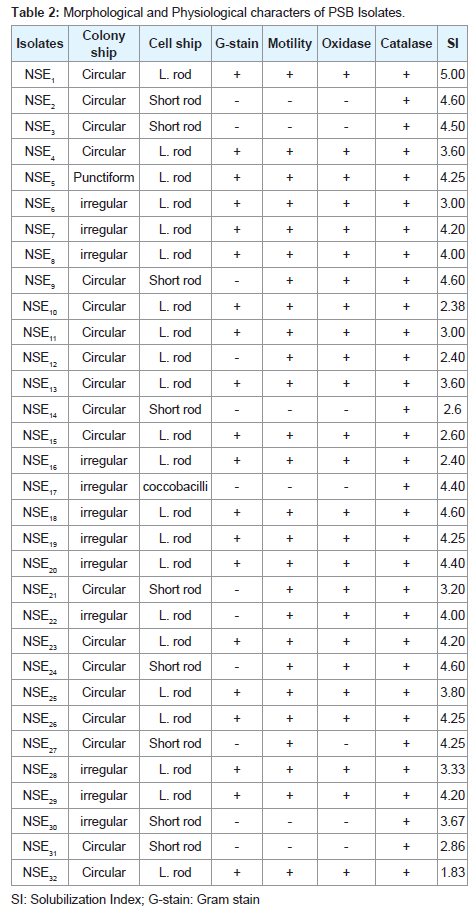
Thirty-two bacterial isolates were obtained from two samples in PKV medium. The possible morphological and physiological characters of the isolates are summarized in (Table 2). The screened bacteria were able to solubilize TCP on solid culture state by forming clear halo zone, with different degree of solubilization, depending on the type of organism involved. However, all the selected isolates were found to be potent phosphate solubilizers showing clear halo zone around their colonies. Among of 32 potent isolates, strain NSE1 showed the maximum phosphate solubilization activity as visualized by the size of clear zone developed around the colony, which showed solubilization index of 5.00 as showed in the same table.
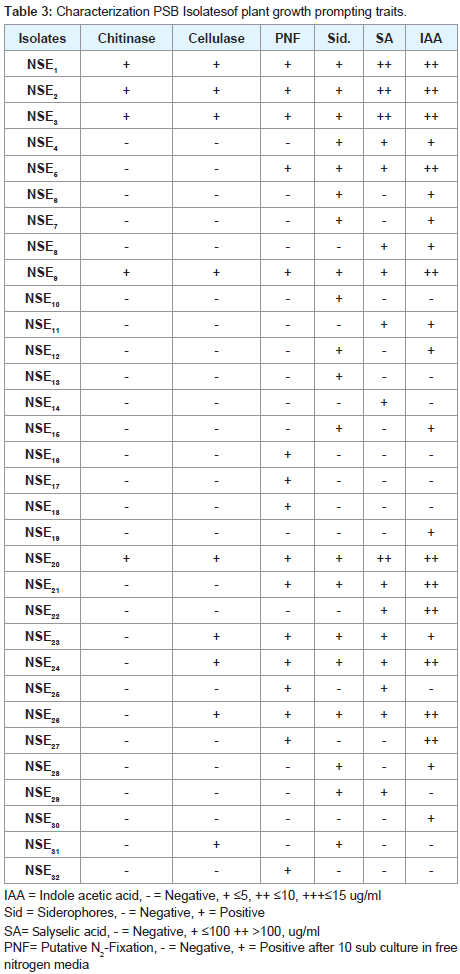
Characterization of plant growth prompting traits
The capability of isolates to produce some plant growth promoting traits was also determined as shown in (Table 3). A total 32 isolates which isolated from different samples tested as plant growth promoters bacteria 21 (68.75%) of them were capable to produce IAA in detectable amounts (5 to 15 μg/ml). Four isolates (12.5%), out of total, produce salicylic acid in appreciable amounts (>100 μg/ml), while 13(40.62%) of them produce less than 100 μg/ml and the rest cannot produce the compound. Concerning Siderophores (Sid), it was found that 20 (62.5%) of the total PSB isolates able to produce Sid. The ability of different isolates to grow on N-free medium (putative N2 fixers) was also examined in suitable liquid culture medium. The isolate capable of growing in medium free of any N-source for ten successive subculture was considered as putative N2 fixer, whether the fixation was via N2-ase or by scavenging nitrogen from the surroundings. In this study, out of 32 isolates tested 16 (50%) of isolates were able to grown on in free medium. Regarding lyticenzymes, among the enzymes tested number of cellulose producers were nine isolates (28.12%).whereas chitinase-producers recorded five isolates (15.62%).
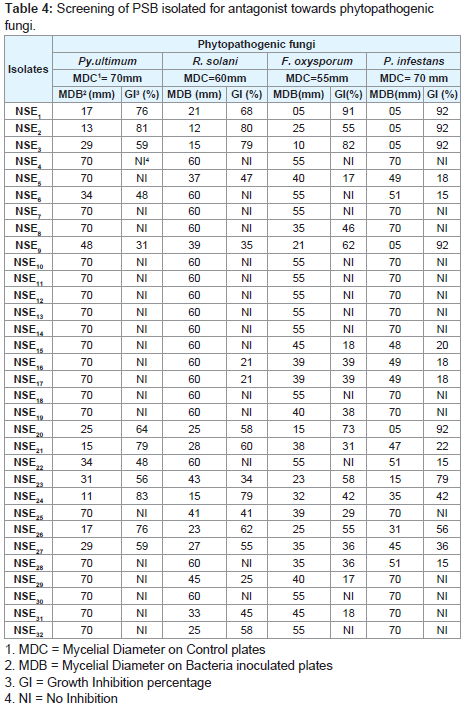
Antifungal activity assay
Plant Growth Promoting Rhizobacteria (PGPR) has attracted the attention of many researchers because of the potential for developing these bacteria as inocula for plant disease control. Results presented in that 12 (37.5%) of tested isolates had a wide range of antagonistic activity against Py. ultimum (Table 4), while about 17 (53.12%), 20 (62.5%) and17 (53.12%), against R. solani F. oxysporum and P. infestans respectively.
Table 4: Representative pictures of Mandukparni (Centella asiatica) callus on day 0 and 7 in control and Biofield Energy Treated groups.
Screening of PSB isolated for antagonist towards phytopathogenic fungi.
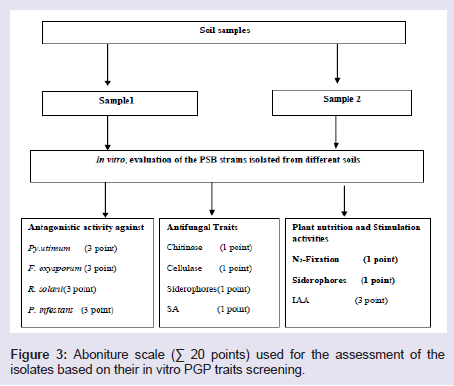
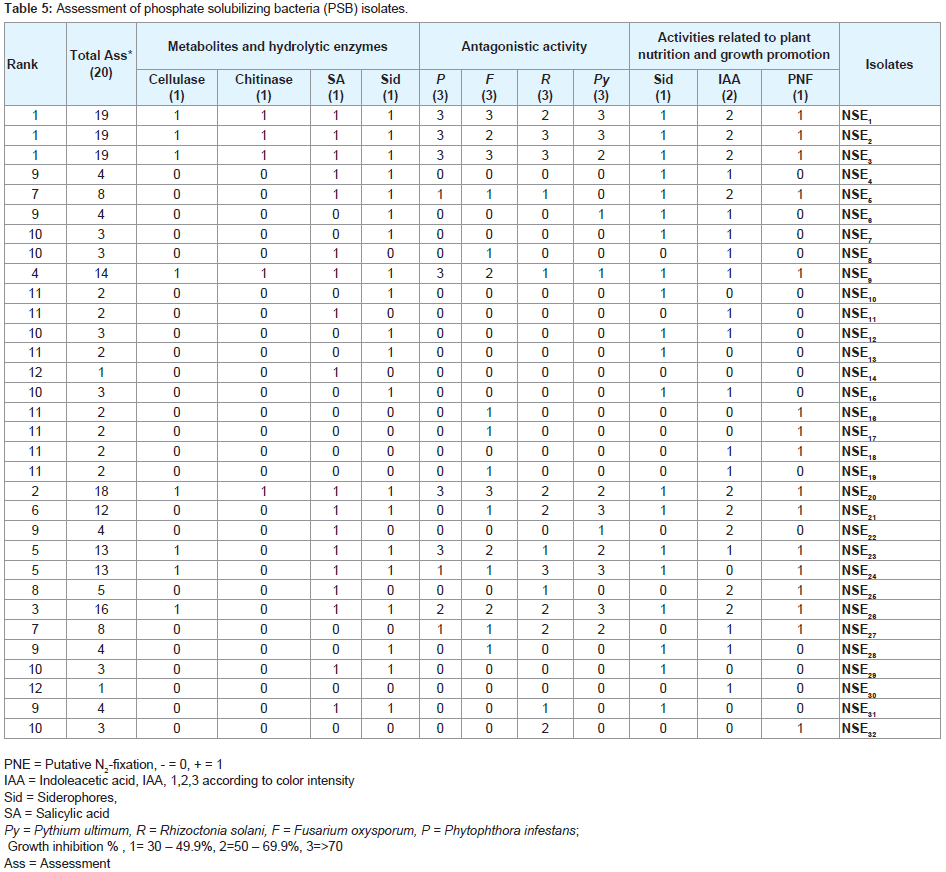
In an attempt to select better bacterial isolates with high plant growth promotion potential, a bonitur scale similar to that described by Krechel was generated and used for assessment of PSB isolates [36] (Figure 3). In this scale, points were given to each bacterial trait in vitro determined within this study. Up to three points each were given for antagonistic activity toward each of the five indicator fungi, one point for each of the hydrolytic enzymes, and PNF, three points for IAA production, and Siderophores were given two points, one as antifungal trait and one for their use by plants for iron acquisition. This generated of bonitur scale of 21 points. As shown in (Table 5), results of the assessment revealed that out of the 32 isolates screened, NSE1, NSE2, NSE3, NSE20 and NSE26 isolates have the top three ranks according to assessment values.
Figure 3: Aboniture scale (∑ 20 points) used for the assessment of the isolates based on their in vitro PGP traits screening.
The five isolates characterized by high efficiency in different activities according to assessment values were tasted effect of pH, temperature and salt content on growth of strains under extreme conditions the results as shown in (Table 6).
The more efficient three strains (NSE1, NSE2 and NSE3) were chosen to study their efficiencies in some important biological processes and activities. The activities studied were limited in N2-fixation ability, cellulose activity, indole acetic acid, siderophores and salicylic acid production. In addition, their inhibition capacity for the growth of four pathogenic fungi which mentioned before. Worth mentioning that the three strains had the same efficiency whether in extreme or in normal conditions.
Molecular characterization and identification of bacterial isolates
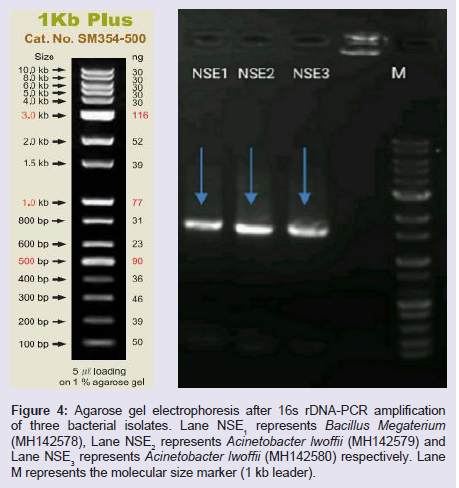
PCR amplification of ribosomal DNA was carried out with universal forward and reverse primers of 16s rDNA and produced a fragment of approximately 1500 bp (Figure 4). This size corresponded to the expected size as compared to other bacteria [37]. The variability within the amplified regions was investigated by phylogenetic analysis. The amplified PCR product was run on1.8% agarose gel viewed under the UV transilluminator.
Figure 4: Agarose gel electrophoresis after 16s rDNA-PCR amplification of three bacterial isolates. Lane NSE1 represents Bacillus Megaterium (MH142578), Lane NSE2 represents Acinetobacter lwoffii (MH142579) and Lane NSE3 represents Acinetobacter lwoffii (MH142580) respectively. Lane M represents the molecular size marker (1 kb leader).
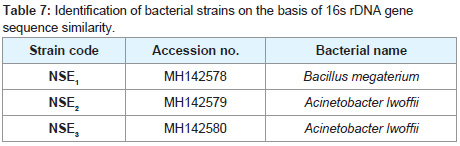
Phylogenetic analysis of the rDNA sequences
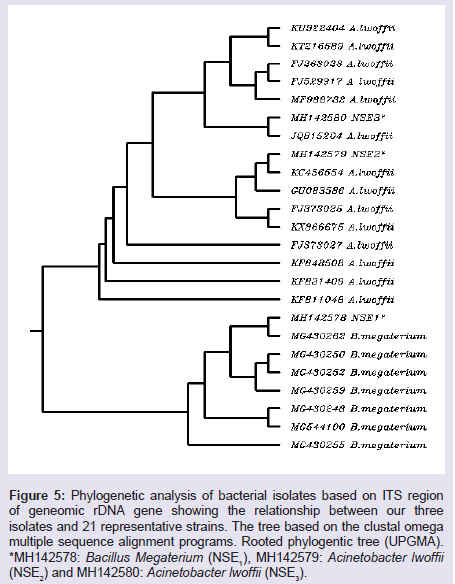
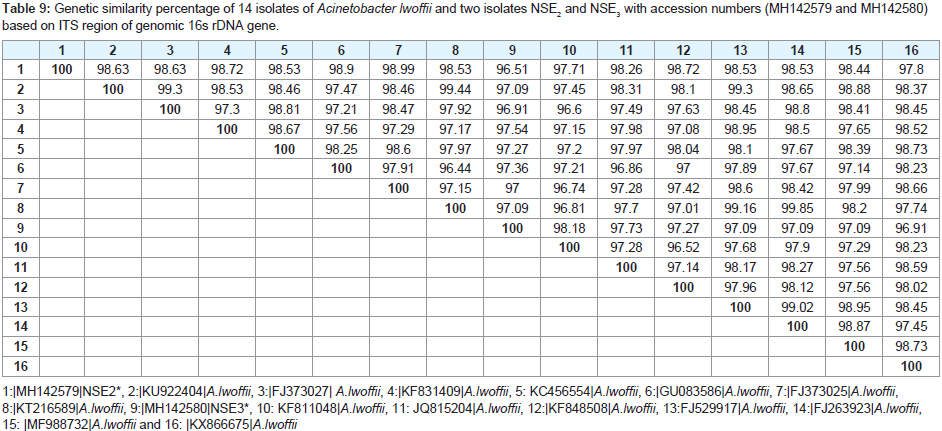
The amplified PCR product of representative isolates was identified and sequenced. A BLASTN analysis carried out through GenBank (http://www.ncbi.nlm.nih.gov) revealed that all the isolates were members of two different genera, Bacillus genus and the genus Acinetobacter as shown in (Table 7). Phylogenetic tree divided into two clusters. The first cluster includes 14 isolates of Acinetobacter lwoffii and two isolates NSE2 and NSE3 with accession numbers (MH142579 and MH142580). The second cluster includes 7 isolates of Bacillus megaterium and our isolate NSE1 with accession number MH142578 as shown in (Figure 5).
Figure 5: Phylogenetic analysis of bacterial isolates based on ITS region of geneomic rDNA gene showing the relationship between our three isolates and 21 representative strains. The tree based on the clustal omega multiple sequence alignment programs. Rooted phylogentic tree (UPGMA). *MH142578: Bacillus Megaterium (NSE1), MH142579: Acinetobacter lwoffii (NSE2) and MH142580: Acinetobacter lwoffii (NSE3).
Discussion
Phosphorus is commonly deficient in most natural soils, since it is fixed as insoluble iron and aluminum phosphates in acidic soils (pH lower than 5.0) or calcium phosphates in alkaline soils (pH above 7.0), as in the Egyptian soils. Phosphate Solubilizing Bacteria (PSB) play a great role in enhance the availability of phosphorus to plants by converting it into soluble form and increasing the crop yield [38,39]. Phosphate Solubilizing Bacteria (PSB) was found in majority of soils, various PSB have been isolated from different plant roots [3,40]. Hence, PSB can be regarded as one kind of plant growth-promoting rhizobacteria, which are widely considered as alternatives to common biofertilizers, also PSB have been found to alleviate harmful impacts of agrochemical [41-44].
In this study 32 strains were isolated as potential phosphate solubilizers based on their ability to solubilize tricalcium phosphate [Ca3 (PO4)2] by formation of a clear zone of solubilization around the colony on Pikovskaya agar medium, Solubilization Index (SI) was determined between 1.83 to 5.00.
The PSB isolates were screened for a wide array of traits that might be associated with ability to function as PGPR. Although many studies have been conducted to identifying the specific traits by which PGPR promote plant growth, usually they were limited to studying just one or two of these traits [45].
In the present work, the 32 PSB isolates were screened for a wide array of traits that might be associated with ability to function as PGPR several bacterial strains have been reported to fix N2. Among of 32 PSB isolated in the present study [45-47], 16 (50%) were found Presumptive Nitrogen Fixers (PNF). Indole Acetic Acid (IAA) is one of the most physiologically active auxins. IAA is a common product of L-tryptophane metabolism by several microorganisms including PGPR [48]. It is presumed that PGPR producing plant growth promoting agents play a critical role in plant growth promotion. In the present work, the 32 PSB isolates were screened for their ability to produce IAA. Results presented in this study show that 68.5% of the PSB isolates produced detectable levels of IAA in culture supernatants. These isolates varied greatly in their ability to produce IAA.
It was reported that the ability of bacterial strains to antagonize pathogenic fungi was related to the production of extracellular siderophores which deprive phytopathogenic microflora of iron, thus limiting their growth [49,50]. It was reported that plants use microbial Sid for iron acquisition [51], and Sid is among factors involved in ISR [52,53]. Concerning Siderophores (Sid), it was found that a 25 % of the total PSB isolates able to produce Sid.
With regard to SA, the data showed that ability to produce SA appears to be widespread among the PSB isolates. More than half of isolates (53%) were able to produce SA. Many studies indicated that SA plays an important role in plant defense response against pathogen attack and is essential for development of both SAR and ISR in plants [54,55].
Biological control of plant pathogens has been the focus of many studies in plant protection that search for alternative or complementary methods to the use of chemical pesticides. PGPR have captured the attention of many researchers because of the potential for developing these bacteria as inocula for plant disease control. The PGPR under most scrutiny for potential use in agriculture are those belonging to the genera Pseudomonas and Bacillus [56]. In the present work, of the 25 isolates showed antifungal activity12 (37.5%) 17 (53.12%), 20 (62.5%) and 17 (53.12%), against Py. ultimum R. solani F. oxysporum P. infestans respectively.
We can summarize the extremophiles and the extremoduric properties in some abbreviations as mentioned by Satyanarayana in the following: Psychrophiles, Thermophiles, Acidophiles, Alkalophiles, Baraophiles, Halophiles, Oligocarophiles, Oligonitrophiles, Radiation resistant, Methanogenic, Toxitolerant, Xerophiles and Organic Solvents Tolerant [57].
In the present study Five strains were chosen because they proved to have the higher activities in the different characters studied (IAA, SA, N2-fixition ...etc) according to assessment values, All isolates were capable of growing at high temperature (50 °C) and pH 7.0 producing observable growth in absence of salt after 24 hours incubation period. This phenomenon was also observed when the pH increased to 10.0 but temperature was at 30 °C in absence of salt but the growth was pronounced after 48 hours, in all strains tested. In presence of 15% salt at pH 10.0 and 50 oC the picture seemed to somewhat different as three strains only were capable of producing observable growth. They were strains No NSE1, NSE2 and NSE3 after 48 hours incubation period. All the three isolates (NSE1, NSE2 and NSE3) showed an inhibitory (in vitro) against the test fungi. The three strains had the same efficiency whether in extreme or in normal conditions.
Molecular studies for the three bacterial isolates with its region of genomic rDNA gene showed band with approximately size 1500 bp using 16s rDNA - PCR two different bacterial genera, Bacillus and Acinetobacter showed closed phylogenetic tree between two isolates of Acinetobacter lwoffii, NSE2 and NSE3 and other subunit with NSE1 from Bacillus megaterium. It means that there are relationships between the two bacterial isolates (Acinetobacter lwoffii, NSE2 and NSE3) and 14 representative strains, and there are relationships between the one bacterial isolate (Bacillus megaterium NSE1) and 7 representative strains.
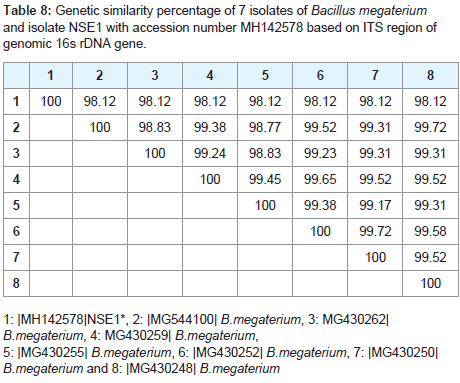
The results of cluster analysis (Similarity index) showed that the highest similarity value 99.44 observed between 2 (KU922404) A. lwoffii and 99.85 between 8 (KT216589) A. lowoffii, 99.85 between 8 KT216589) A. lwoffii and 14 (Fj263923) A.lwoffii. The Lowest similarity was 96.51 between 1 (MH142579) NSE2 and 9 (MNH142580) as shown in (Table 8 and 9).
Table 8: Genetic similarity percentage of 7 isolates of Bacillus megaterium and isolate NSE1 with accession number MH142578 based on ITS region of genomic 16s rDNA gene.
Conclusions
Considerable research efforts are underway globally to exploit the potential of Phosphate Solubilizing Bacteria (PSB) as alternative for chemical pesticides and chemical fertilizers. The main objective of this thesis was to find strains of Phosphate Solubilizing Bacteria (PSB), have broad spectrum of plant growth-promoting abilities and antagonistic potential against phytopathogenic fungi, which could be used as safe alternative for the overuse of harmful agrochemicals. In general, The most three effective isolates were identified as Bacillus megaterium MH142578, Acinetobacter lwoffii MH142579 and Acinetobacter lwoffii MH142580 which have plant growth-promoting abilities and antagonistic potential against phytopathogenic fungi involved on anther proceeding study.
References
- Pradhan N and Sukla LB (2005) Solubilization of inorganic phosphate by fungi isolated from agriculture soil. African J Biotechnol 5.
- Guiñazú LB, Andrés JA, Del Papa MF, Pistorio M, Rosas SB (2010) Response of alfalfa (Medicago sativa L.) to single and mixed inoculation with phosphate-solubilizing bacteria and Sinorhizobium meliloti. Biol Fert Soils 46: 185-190.
- Yu X, Liu X, Zhu T, Liu G, Mao C (2011) Isolation and characterization of phosphate-solubilizing bacteria from walnut and their effect on growth and phosphorus mobilization. Biol Fert Soils 47: 437-446.
- Khosro Mohammadi, Yousef Sohrabi (2012) Bacterial biofertilizers for sustainable crop production: A REVIEW. ARPN J Agr Biological Sci 7: 307-315.
- Gyaneshwar P, Naresh Kumar G, Parekh LJ, Poole PS (2002) Role of soil microorganisms in improving P nutrition of plants. Plant and Soil 245: 83-93.
- Hao X, Cho CM, Racz GJ, Chang C (2002) Chemical retardation of phosphate diffusion in an acid soil as affected by liming. Nutr Cycl Agroecosys 64: 213-224.
- Shahid M, Khan MS (2017) Assessment of glyphosate and quizalofop mediated toxicity to greengram [Vigna radiate (L.) Wilczek], stress abatement and growth promotion by herbicide tolerant Bradyrhizobium and Pseudomonasspecies. Int J Curr Microbiol App Sci 6: 3000-3001.
- Richardson AE (2001) Prospects for using soil microorganisms to improve the acquisition of phosphate solubilization. 69-75.
- Alam SS, Ayub KN, Rashid M (2002) In vitro solubilization of inorganic phosphate by phosphate solubilizing microorganism (PSM) from maize rhizosphere. Intl J Agric Biol 4: 454-458.
- Antoun H, Kloepper JW (2001) Plant growth-promoting rhizobacteria (PGPR), In: Encyclopedia of Genetics, Brenner S, Miller JH (Eds) Academic Press, N.Y., pp.1477-1480.
- Klute A (1986) Methods of Soil Analysis. Part-1: Physical and Mineralogical Methods. 2nd Edition. American Society of Agronomy, Madison, Wisconsin, USA.
- Page AL, Miller RH, Keeney DR (1982) Methods of Soil Analysis. Part-1: Chemical and Microbiological Properties. 2nd Edition. American Society of Agronomy, Madison, Wisconsin, USA.
- Chapman HD, Pratt PF (1961) Method of analysis for soils and water. University of California, Div. Agric. Sci., USA.
- Jackson ML (1956) Soil analysis. Adv. Course. Fourth print. Dept. of Soil Sci. Univ. of Wisconsin, Madison. WI.
- Black CA, Evans DD, White JL, Ensminger LE, Clark FE (1965) Methods of soil analysis" part 1: physical and mineralogical properties. Published by the American Society of Agronomy. Madison, Wisconsin, USA.
- Olsen SR, Sommers LE (1982) Phosphorus In: Page AL, Miller RH and Keeney DR. (Eds.). Methods of Soil Analysis. Part 2. 2nd ed. ASA and SSSA Publisher, Madison. 403-430.
- Lindsay WL, Norvell WA (1978) Development of a DTPA soil test for zinc, iron, manganese and copper. Soil Sci Soc Am J 42: 421-428.
- Pikovskaya RI (1948) Mobilization of phosphates in soil in connection with the vital activities of some microbial species. Microbiology 17: 362-370.
- Defreitas JR, Banerjee MR, Gemida JJ (1997) Phosphate solubilizing rhizobacteria enhance the growth and yield but not phosphorus uptake of canola (Brassica napus L.). Biol Fertil Soils 24: 358-364.
- Edi-Premono M, Moawad MA, Vleck PLG (1996) Effect of phosphate solubilizing Pseudmonasputida on the growth of maize and its survival in the rhizosphere. Indonesian J Crop Sci 11: 13-23.
- Loper JE, Schroth MN (1986) Influence of bacterial sources of indole-3-acetic acid on root elongation of sugar beet. Phytopathology 76: 386-389.
- Schwyn B, Neilands JB (1987) Universal chemical assay for detection and determination of siderophores. Anal Biochem 160: 47- 56.
- Pallai R (2005) Effect of plant growth-promoting rhizobacteria on canola (Brassica napus. L) and lentil (Lens culinaris. Medik) plants. Thesis, Master of Science, Department of Applied Microbiology and Food Science, University of Saskatchewan, Saskatoon. Canada.
- Meyer JM, Abdallah MA (1978) The fluorescent pigment of Pseudomonas fluorescens: biosynthesis, purification and physicochemical properties. Journal of General Microbiology 10: 319-328.
- Leeman M, Denouden FM, van Pelt JA, Cornelissen C, Matamala Garros A, et al. (1996) Suppression of Fusarium wilt of radish by co-inoculation of fluorescent Pseudomonas spp. and root-colonizing fungi. Eur J Plant Pathol 102: 21-31.
- Andro T, Chambost JP, Kotoujansky A, Cattaneo J, Bertheau Y, et al. (1984) Mutants of Erwiniachrysanthemi defective in secretion of pectinase and cellulase. Journal of Bacteriol 160: 1199-1203.
- Frändberg E, Schnürer J (1998) Antifungal activity of chitinolytic bacteria isolated from airtight stored cereal grain. Can J Microbiol 44: 121-127.
- Haahtela K, Helander I, Nurmiaha-Lassila EL, Sundman V (1983a) Morphological and physiological characteristics and lipopolysaccharide composition of N2-fixing (C2H2-reducing) root-associated Pseudomonas sp. Can J Microbiol 29: 874-880.
- Haahtela K, Kari K, Sundman V (1983b) Nitrogenase activity (Acetylene Reduction) of root-associated, cold- climate Azosperillum, Enterobacter, Klabsilla, and Pseudomonas species. Appl Environ Microbiol 45: 563-570.
- Cattelan AJ, Hartel PG, Fuhramnn JJ (1999) Screening for plant growth-promoting to promote early soybean growth. Soil Sci Soc Am J 63: 1670-1680.
- Koch E (1997) Screening of rhizobacteria for antagonistic activity against Pythiumultimum on cucumber and kale. J Plant Dis Prot 104: 353-361.
- Miller CS, Handley KM, Wrighton KC, Frischkorn KR, Thomas BC, et al. (2013) Short-read assembly of full-length 16S amplicons reveals bacterial diversity in subsurface sediments. PloS one 8: e56018.
- 33.Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 3389-3402.
- Benson DA, Boguski MS, Lipman DJ, Ostell J, Oullette BF, et al. (1999) GenBank. Nucleic Acids Res 8: 12-17.
- Thompson JD, Higgins DJ, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positive specific gap penalties and weight matrix choice. Nucleic acids Res 22: 4673-4688.
- Krechel A, Faupel A, Hallmann J, Ulrich A, Berg G (2002) Potato-associated bacteria and their antagonistic potential towards plant-pathogenic fungi and the plant-parasitic nematode Meloidogyne incognita (Kofoid and White) Chitwood. Can J Microbiol 48: 772-786.
- Cheng K, Lu FP, Li M, Liu LL, Liang XM (2010) Purification and biochemical characterization of a serine alkaline protease TC4 from a new isolated Bacillus alcalophilus TCCC11004 in detergent formulations. African J Biotechnol 9.
- 38.Jones DL, Darrah PR (1994) Role of root derived organic acids in the mobilization of nutrients from the rhizosphere. Plant Soil 166: 247-257.
- Toro M, Azcon R, Barea JM (1997) Improvement of arbuscularmycorrhiza development by inoculation of soil with phosphate-solubilizing rhizobacteria to improve rock phosphate bioavailability (sup32) P and nutrient cycling. Appl Environ Microbiol 63: 4408-4412.
- Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 3389-3402.
- Vessey JK (2003) Plant growth promoting rhizobacteria as biofertilizers. Plant and Soil 255: 571-586.
- Hafeez FY, Yasmin S, Ariani D, Mehboob UR, Zafar RY, et al. (2006) Plant growth-promoting bacteria as biofertilizer. Agron Sust Develop 26: 143-150.
- Shahid M, Ahmed B, Khan MS (2018) Evaluation of microbiological management strategy of herbicide toxicity to greengram plants. Biocatalysis and Agricultural Biotechnology 14: 96-108.
- Adesemoye AO, Torbert HA, Kloepper JW (2009) Plant growth-promoting rhizobacteria allow reduced application rates of chemical fertilizers. Microbial Ecol 58: 921-929.
- Cattelan AJ, Hartel PG, Fuhramnn JJ (1999) Screening for plant growth-promoting to promote early soybean growth. Soil Science Society of America Journal 63: 1670-1680.
- Chen YP, Rekha PD, Arunshen AB, Lai WA, Young CC (2006) Phosphate solubilizing bacteria from subtropical soil and their tricalcium phosphate solubilizing abilities. Appl Soil Ecol 34: 33-41.
- Andrade G, Esteban E, Velasco L, Lortie MJ, Bedmar EJ (1997) Isolation and identification of N2-fixing microorganisms from the rhizoplane of Capparisspinosa (L.). Plant and Soil 197: 19-23.
- El-Sayed WS, Akhkha A, El-Naggar MY, Elbadry M (2014) Invitroantagonisticactivity plant growth promoting traits and phylogenetic affiliation of rhizobacteria associated with wild plants grown in arid soil. Front Microbiol 5: 1-11.
- Kloepper JW, Leong J, Teintze M, Schroth MN (1980b) Pseudomonas siderophores: A mechanism explaining disease-suppressive soils. Current Microbiology 4: 317-320.
- Leong J (1986) Siderophores: their biochemistry and possible role in the biocontrol of plant pathogens. Ann Rev Phytopathol 26: 187-209.
- Yehuda Z, Shenker M, Romheld V, Marschner H, Hadar Y, et al. (1996) The role of legend exchange in the Uptake of Iron from microbial siderophores by gramineous plants. Plant Physiol 112: 1273-1280.
- Leeman M, van Pelt JA, Hendrickx MJ, Scheffer RJ, Bakker PAHM (1995a.) Biocontrol of fusarium wilt of radish in commercial greenhouse trials by seed treatment with Pseudomonas fluorescens WCS374. J Phytopathology 85: 1301-1305.
- De Mayer G, Höfte M (1997) Salicylic Acid Produced by the rhizobacterium Pseudomonas aeruginosa 7NSK2 induces resistance to leaf Infection by Botrytis cinerea on bean. Phytopathology 86: 588-593.
- van Loon LC, Bakker PA, Pieterse CM (1998) Systemic Resistance Induced by rhizosphere bacteria. Annu Rev Phytopathol 36: 453-483.
- Zhang S, Moyne L, Reddy MS, Kloepper JW (2002) The role of salicylic acid in induced systemic resistance elicited by plant growth-promoting rhizobacteria against blue mold of tobacco. Biocontrol 25: 288-296.
- Powell KA, Rhodes DJ (1994) Strategies for the progression of biological fungicides into field evaluation. British Crop Protection Council Symposium Proceedings, BCPC Monograph. No. 59: 307-315.
- Satyanarayana T, Raghukumar C, Shivaji S (2005) Extremophilic microbes: Diversity and perspectives. Curr Sci 89: 78-90.