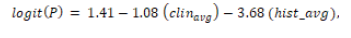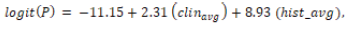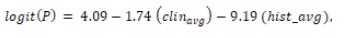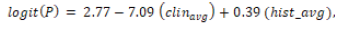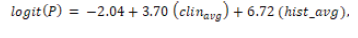Journal of Clinical and Investigative Dermatology
Download PDF
It comprises data from 366 patients, each diagnosed with one of six erythemato-squamous skin conditions: psoriasis, seborrheic dermatitis, lichen planus, pityriasis rosea, chronic dermatitis, or pityriasis rubra pilaris. Each patient record includes a total of 34 attributes, of which 33 are feature variables, and 1 is the target class label. This attribute is coded as an integer from 1 to 6, corresponding to the six disease types:
• Scaling
• Itching
• Definite borders
• Koebner phenomenon
• Polygonal papules
• Follicular papules
• Oral mucosal involvement
• Knee and elbow involvement
• Scalp involvement
• Family history (binary: 0 = no family history, 1 = positive family history)
• Age (continuous, with eight missing values)
These clinical markers form the basis of initial differential diagnosis and reflect the physician’s visual and tactile interpretation of disease. These are also the more obvious and irritating features that have direct effects on patients’ lives.
• Melanin incontinence
• Eosinophils in the infiltrate
• PNL infiltrate
• Fibrosis of the papillary dermis
• Exocytosis
• Acanthosis
• Hyperkeratosis
• Parakeratosis
• Clubbing of the rete ridges
• Elongation of the rete ridges
• Thinning of the suprapapillary epidermis
• Spongiform pustule
• Munro microabscess
• Focal hypergranulosis
• Disappearance of the granular layer
• Vacuolization and damage of basal layer
• Spongiosis
• Saw-tooth appearance of rete ridges
• Follicular horn plug
• Perifollicular parakeratosis
• Inflammatory mononuclear infiltrate
• Band-like infiltrate These features provide deeper insight into the structural and inflammatory processes and are commonly used to confirm clinical suspicion when overlap is high.
One of the earliest landmark studies conducted by Güvenir et al. [1] first utilized the Dermatology dataset. They introduced a rule-based classification model, the Voting Feature Intervals (VFI5) algorithm, and laid the foundation for interpreting hybrid diagnostic models. While it did not directly compare the predictive strength of clinical versus histopathological data, it established the value of using both feature sets for optimal classification accuracy.
As the field progressed, Übeyli and Doğdu [13] applied unsupervised learning through k-means clustering to the same dataset. This method grouped patients based on feature similarity and disregarded prior class labels. They demonstrated that the data held clear discriminatory power, despite being unsupervised. Additionally, they showed that histopathological features had a slightly stronger influence on cluster formation.
Other studies introduced a classification and regression tree (CART) method of modeling and applied it to this dataset.[14,15] As one of the earlier applications of decision tree modeling, CART proved to be a practical and interpretable tool. The model did well and outperformed more complex neural networks at the time, reinforcing the strength of straightforward, rule-based classifiers.
Elsayad et. al. [16] expanded on this decision tree methodology using the CHAID (Chi-Squared Automatic Interaction Detection) model. This approach allowed for multi-level branching based on statistical associations between variables and disease class. They also addressed algorithm instability by implementing bagging and boosting models to enhance model robustness while retaining interpretability.
After importing the dataset using the readxl package in R, clinical and histopathological features were grouped into two respective categories based on domain knowledge. The rowMeans() function was used to compute the average clinical and pathological scores per patient. From these averages, we generated composite predictors representing overall clinical and histopathological severity for each subject.
We performed a paired t-test using these averages to determine whether clinical and histopathological measurements differed. In addition, patients were stratified by their class label, indicating one of the six ESDs, and separate paired t-tests were conducted within each diagnostic group. The paired structure accounted for repeated measurements on the same individuals. Both histograms and boxplots indicate that the differences approximately follow a normal distribution. Our analysis of individual disease classes helped assess whether certain diseases rely more heavily on clinical or pathological characteristics for accurate classification.
Other graphical tools were implemented to supplement the statistical analyses and assess underlying assumptions. As mentioned previously, boxplots and histograms were created as summary visualizations to assess symmetry and potential deviations from normality, like skewness and outliers. Back-to-back boxplots helped compare the two groups of data and their distributions by class. We also used paired line plots as an extra method to visually assess differences within each class. These visualization techniques were not only used for exploratory purposes but also for validation of paired t-test model assumptions.
We fit six multiple logistic regression models to assess the relative predictive power of clinical and histopathological features across each ESD. For each model, the outcome variable was binary (1 = presence of a specific ESD class/disease; 0 = all other classes/diseases). The model predictors included the average clinical score and average pathological score across the respective feature sets. The models were built using the glm() function in R with a binomial family and logit link. This approach enabled the estimation of the independent contribution of clinical and pathological data in classifying disease presence, while also allowing for direct comparison of predictive trends across disease types.
All analyses were conducted using R version 4.3.1. In addition to readxl, packages utilized in this analysis include dplyr for data wrangling, ggplot2 for data visualization, tidyr for reshaping data, and ggpubr for graphical hypothesis testing annotation. These tools enabled reproducible, transparent, and efficient statistical modeling across the dataset as a whole and individual disease classes
Research Article
Comparing Clinical vs Histopathological Features in Diagnosing Erythemato-Squamous Diseases
Shenouda MG1,2, Travers JB1,2,3* and Sun S4
1Wright State University Boonshoft School of Medicine, Dayton, Ohio, USA
2Departments of Pharmacology and Toxicology, Wright State University, Dayton, Ohio, USA
3Department of Dermatology, Wright State University, Dayton, Ohio, USA
4Department of Mathematics and Statistics, Wright State University, Dayton, Ohio, USA
2Departments of Pharmacology and Toxicology, Wright State University, Dayton, Ohio, USA
3Department of Dermatology, Wright State University, Dayton, Ohio, USA
4Department of Mathematics and Statistics, Wright State University, Dayton, Ohio, USA
*Address for Correspondence:Jeffrey B. Travers, Wright State University Department of
Pharmacology and Toxicology, 3640 Colonel Glenn Hwy, Dayton, OHIO. USA. E-mail Id: jeffrey.travers@wright.edu
Submission: 21 February, 2026
Accepted: 24 March, 2026
Published: 26 March, 2026
Copyright: ©2026 Shenouda MG, et al. This is an open access
article distributed under the Creative Commons Attribution License,
which permits unrestricted use, distribution, and reproduction in any
medium, provided the original work is properly cited.
Keywords:Erythemato-squamous diseases; Clinical features; Histopathology;
Diagnostic classification; UCI Dermatology dataset; Logistic regression; Predictive accuracy
Abstract
Erythemato-squamous diseases (ESDs) possess overlapping clinical
manifestations and diverse histopathological profiles, thus presenting
diagnostic challenges. There remains a need for improved diagnostic
approaches that integrate clinical and histopathological features.
The objective of these studies is to investigate the relative value of
clinical versus histopathological features in distinguishing among
six ESD classes. The University of California, Irvine (UCI) Dermatology
dataset includes 366 patients diagnosed with one of six ESDs and
their corresponding clinical and histopathological features. Data
were analyzed using paired t-tests. Multiple logistic regression (MLR)
models were constructed for each ESD class to assess the predictive
strength of clinical and histopathological features. Paired t-tests
revealed a statistically significant difference between clinical and
histopathological averages across the dataset (p < 2.2e-16), with
clinical features generally more pronounced. This trend was consistent
across all disease classes except chronic dermatitis, where no significant
difference was observed (p = 0.8102). Multiple logistic regression
models demonstrated high predictive performance across all six ESD
classes, with pityriasis rubra pilaris achieving the highest predictive
accuracy of 94.5%. Clinical features exhibited higher average severity
across the dataset; however, this does not necessarily translate into
diagnostic dominance, which varies by disease class. For conditions
like lichen planus, histopathological features provided stronger
predictive power. Our results underscore the complementary roles of
clinical and histopathological data and support the development of
integrated models for improving classification accuracy and data driven
diagnostic strategies in dermatology.
Abbreviations
ESD: Erythemato-squamous skin diseases; GrC: Granular
computing; kNN: K-nearest neighbors; LP: Lichen planus; PR:
Pityriasis rosea; PRP: Pityriasis rubra pilaris; SVC: Support vector
classifier; SVM: Support vector machines.
Introduction
Erythemato-squamous skin diseases (ESDs) have been classified
to involve 6 different skin conditions that show very close findings
upon clinical examination. These six have been classified to include
psoriasis, seborrheic dermatitis, lichen planus, pityriasis rosea,
chronic dermatitis, and pityriasis rubra pilaris.[1,2] Similarities, such
as erythema and scaling, are present in almost all of these conditions,
with very few differences. Such chronic diseases take a toll on the
mental, emotional, and physical stress of patients.
Therefore, prompt and accurate diagnosis and treatment of these diseases is critical. However, not only do these diseases share many clinical features, but they are often similar in histopathological features as well. In the field of dermatology, histopathological evaluation is treated as the gold standard for the diagnosis of skin diseases.[3-5] As such, a biopsy is usually required for diagnosis; however, its effectiveness and clinical utility remain debated. Overlapping clinical and histopathological features complicate accurate diagnosis, particularly when differentiating between premalignant and malignant lesions.[6-10] Low concordance between clinical and histopathological assessments can compromise correct diagnosis and harm patient care.[11] Comparing the predictive power of these variables is central to ongoing discussions in this field. This study aimed to compare the diagnostic contribution of clinical and histopathological features in differentiating six erythematosquamous diseases using statistical methods and predictive modeling applied to the UCI Dermatology dataset.
Therefore, prompt and accurate diagnosis and treatment of these diseases is critical. However, not only do these diseases share many clinical features, but they are often similar in histopathological features as well. In the field of dermatology, histopathological evaluation is treated as the gold standard for the diagnosis of skin diseases.[3-5] As such, a biopsy is usually required for diagnosis; however, its effectiveness and clinical utility remain debated. Overlapping clinical and histopathological features complicate accurate diagnosis, particularly when differentiating between premalignant and malignant lesions.[6-10] Low concordance between clinical and histopathological assessments can compromise correct diagnosis and harm patient care.[11] Comparing the predictive power of these variables is central to ongoing discussions in this field. This study aimed to compare the diagnostic contribution of clinical and histopathological features in differentiating six erythematosquamous diseases using statistical methods and predictive modeling applied to the UCI Dermatology dataset.
Methods
Data Source:
Due to the complexity of diagnosing ESDs, research has been
productive in determining what models best convey the most accurate
predictive diagnoses based on clinical and histopathological findings.
To support this research, a publicly available resource known as the
“Dermatology dataset” was donated by Güvenir et al. (1998) [1] and
is now hosted by the University of California, Irvine (UCI) Machine
Learning Repository.[12] It has been used extensively for machine
learning and diagnostic classification research in dermatology. This
dataset enables comparative analyses of various predictive models in
distinguishing among the 6 ESDs.It comprises data from 366 patients, each diagnosed with one of six erythemato-squamous skin conditions: psoriasis, seborrheic dermatitis, lichen planus, pityriasis rosea, chronic dermatitis, or pityriasis rubra pilaris. Each patient record includes a total of 34 attributes, of which 33 are feature variables, and 1 is the target class label. This attribute is coded as an integer from 1 to 6, corresponding to the six disease types:
• 1 = Psoriasis (n =112)
• 2 = Seborrheic dermatitis (n = 61)
• 3 = Lichen planus (n = 72)
• 4 = Pityriasis rosea (n = 49)
• 5 = Chronic dermatitis (n = 52)
• 6 = Pityriasis rubra pilaris (n = 20)
• 2 = Seborrheic dermatitis (n = 61)
• 3 = Lichen planus (n = 72)
• 4 = Pityriasis rosea (n = 49)
• 5 = Chronic dermatitis (n = 52)
• 6 = Pityriasis rubra pilaris (n = 20)
Clinical Features of Disease:
This dataset includes 12 clinical attributes derived from physical
examination. Each is rated on an ordinal scale from 0 (absent) to 3
(maximal presence) unless otherwise noted. These features include:
• Erythema• Scaling
• Itching
• Definite borders
• Koebner phenomenon
• Polygonal papules
• Follicular papules
• Oral mucosal involvement
• Knee and elbow involvement
• Scalp involvement
• Family history (binary: 0 = no family history, 1 = positive family history)
• Age (continuous, with eight missing values)
These clinical markers form the basis of initial differential diagnosis and reflect the physician’s visual and tactile interpretation of disease. These are also the more obvious and irritating features that have direct effects on patients’ lives.
Histopathological Features of Disease:
This dataset also includes 22 histopathological features of ESDs.
These are each derived from microscopic examination of skin
biopsies. Similar to the clinical attributes, each histopathological
feature is rated on an ordinal scale from 0 (absent) to 3 (maximal
presence). These include:• Melanin incontinence
• Eosinophils in the infiltrate
• PNL infiltrate
• Fibrosis of the papillary dermis
• Exocytosis
• Acanthosis
• Hyperkeratosis
• Parakeratosis
• Clubbing of the rete ridges
• Elongation of the rete ridges
• Thinning of the suprapapillary epidermis
• Spongiform pustule
• Munro microabscess
• Focal hypergranulosis
• Disappearance of the granular layer
• Vacuolization and damage of basal layer
• Spongiosis
• Saw-tooth appearance of rete ridges
• Follicular horn plug
• Perifollicular parakeratosis
• Inflammatory mononuclear infiltrate
• Band-like infiltrate These features provide deeper insight into the structural and inflammatory processes and are commonly used to confirm clinical suspicion when overlap is high.
Study Design:
A wide range of machine learning and statistical methods have
already been applied to the Dermatology dataset to improve the
accuracy of diagnosing ESDs. These approaches range from classic
statistical approaches to complex artificial intelligence (AI)-driven
models, reflecting the growing interest in computational methods for
dermatological diagnostics.One of the earliest landmark studies conducted by Güvenir et al. [1] first utilized the Dermatology dataset. They introduced a rule-based classification model, the Voting Feature Intervals (VFI5) algorithm, and laid the foundation for interpreting hybrid diagnostic models. While it did not directly compare the predictive strength of clinical versus histopathological data, it established the value of using both feature sets for optimal classification accuracy.
As the field progressed, Übeyli and Doğdu [13] applied unsupervised learning through k-means clustering to the same dataset. This method grouped patients based on feature similarity and disregarded prior class labels. They demonstrated that the data held clear discriminatory power, despite being unsupervised. Additionally, they showed that histopathological features had a slightly stronger influence on cluster formation.
Other studies introduced a classification and regression tree (CART) method of modeling and applied it to this dataset.[14,15] As one of the earlier applications of decision tree modeling, CART proved to be a practical and interpretable tool. The model did well and outperformed more complex neural networks at the time, reinforcing the strength of straightforward, rule-based classifiers.
Elsayad et. al. [16] expanded on this decision tree methodology using the CHAID (Chi-Squared Automatic Interaction Detection) model. This approach allowed for multi-level branching based on statistical associations between variables and disease class. They also addressed algorithm instability by implementing bagging and boosting models to enhance model robustness while retaining interpretability.
More recent studies have shifted toward ensemble and hybrid
machine learning models. Bozok and Çalhan [17] compared several
supervised learning methods, including logistic regression, K-nearest
neighbors (kNN), support vector classifier (SVC), Gaussian naïve
Bayes, decision tree, and random forest. Their study found that naïve
Bayes produced up to 100% classification accuracy when used with
histopathological features. Wang and Xie used hybrid methods such
as granular computing (GrC) and support vector machines (SVM)
to draw connections between this dataset and predictive ability.[18]
The increasing complexity of newer algorithms and hybrid models
has fueled even more research in the field.
Recently, Swain et. al. employed a hybrid ensemble model incorporating SVM, logistic regression, kNN, and decision tree classifiers.[19] Their approach involved one-way ANOVA and Chi-square tests for feature selection and GridSearchCV for hyperparameter optimization. They achieved 98.9% accuracy and further reinforced the superiority of combined feature use.
Introducing a more AI-centered approach, a 2025 study conducted by Balbal [20] used a wrapper-based feature selection and compared six machine learning algorithms, again finding naïve Bayes to be the top performer with an accuracy rate of 99.45%. Most notably, this study demonstrated that even reduced feature sets involving fewer than 20 variables could still yield accuracies above 99%. This very recent research continues to affirm the ongoing interest in optimizing diagnostic tools for clinical efficiency without compromising accuracy.
Complementing these modeling advances, Jogu [21] provides a broad review of current machine learning practices in ESD diagnosis. His work highlights the trend towards lightweight, explainable models that balance performance with real-world usability. These types of models help achieve the ultimate goal of providing improved quality care to patients with ESDs.
Finally, Sopjani et. al.[22] contribute a clinical perspective by reviewing agreement rates between clinical and histopathological diagnoses. They reviewed 29 studies where different skin diseases were investigated and assessed the clinico-pathological agreement rates. Their conclusions suggest that while histopathology remains dominant, clinical features alone may offer more value than previously assumed.
As noted above, the field of dermatology has experienced a notable increase in research aimed at improving the diagnosis and management of ESDs. As discovered by the improving predictive ability of models, diagnosing ESDs is largely considered a datamining problem.[23,24] Data mining is a complex means by which valuable information is obtained from large amounts of data. This process draws on multidisciplinary fields such as statistics, artificial intelligence, and machine learning, utilizing a variety of models and algorithms that yield diverse outcomes.[25]
More recent studies have shifted toward ensemble and hybrid machine learning models. Bozok and Çalhan [17] compared several supervised learning methods, including logistic regression, K-nearest neighbors (kNN), support vector classifier (SVC), Gaussian naïve Bayes, decision tree, and random forest. Their study found that naïve Bayes produced up to 100% classification accuracy when used with histopathological features. Wang and Xie used hybrid methods such as granular computing (GrC) and support vector machines (SVM) to draw connections between this dataset and predictive ability.[18] The increasing complexity of newer algorithms and hybrid models has fueled even more research in the field. Recently, Swain et. al. employed a hybrid ensemble model incorporating SVM, logistic regression, kNN, and decision tree classifiers.[19] Their approach involved one-way ANOVA and Chi-square tests for feature selection and GridSearchCV for hyperparameter optimization. They achieved 98.9% accuracy and further reinforced the superiority of combined feature use. Introducing a more AI-centered approach, a 2025 study conducted by Balbal [20] used a wrapper-based feature selection and compared six machine learning algorithms, again finding naïve Bayes to be the top performer with an accuracy rate of 99.45%. Most notably, this study demonstrated that even reduced feature sets involving fewer than 20 variables could still yield accuracies above 99%. This very recent research continues to affirm the ongoing interest in optimizing diagnostic tools for clinical efficiency without compromising accuracy. Complementing these modeling advances, Jogu [21] provides a broad review of current machine learning practices in ESD diagnosis. His work highlights the trend towards lightweight, explainable models that balance performance with real-world usability. These types of models help achieve the ultimate goal of providing improved quality care to patients with ESDs. Finally, Sopjani et. al.[22] contribute a clinical perspective by reviewing agreement rates between clinical and histopathological diagnoses. They reviewed 29 studies where different skin diseases were investigated and assessed the clinico-pathological agreement rates. Their conclusions suggest that while histopathology remains dominant, clinical features alone may offer more value than previously assumed. As noted above, the field of dermatology has experienced a notable increase in research aimed at improving the diagnosis and management of ESDs. As discovered by the improving predictive ability of models, diagnosing ESDs is largely considered a datamining problem.[23,24] Data mining is a complex means by which valuable information is obtained from large amounts of data. This process draws on multidisciplinary fields such as statistics, artificial intelligence, and machine learning, utilizing a variety of models and algorithms that yield diverse outcomes.[25] Despite the increasing complexity of modern analytical approaches, a consistent emphasis remains on balancing accuracy, efficiency, and clinical relevance across the UCI Dermatology dataset and similar datasets.
Recently, Swain et. al. employed a hybrid ensemble model incorporating SVM, logistic regression, kNN, and decision tree classifiers.[19] Their approach involved one-way ANOVA and Chi-square tests for feature selection and GridSearchCV for hyperparameter optimization. They achieved 98.9% accuracy and further reinforced the superiority of combined feature use.
Introducing a more AI-centered approach, a 2025 study conducted by Balbal [20] used a wrapper-based feature selection and compared six machine learning algorithms, again finding naïve Bayes to be the top performer with an accuracy rate of 99.45%. Most notably, this study demonstrated that even reduced feature sets involving fewer than 20 variables could still yield accuracies above 99%. This very recent research continues to affirm the ongoing interest in optimizing diagnostic tools for clinical efficiency without compromising accuracy.
Complementing these modeling advances, Jogu [21] provides a broad review of current machine learning practices in ESD diagnosis. His work highlights the trend towards lightweight, explainable models that balance performance with real-world usability. These types of models help achieve the ultimate goal of providing improved quality care to patients with ESDs.
Finally, Sopjani et. al.[22] contribute a clinical perspective by reviewing agreement rates between clinical and histopathological diagnoses. They reviewed 29 studies where different skin diseases were investigated and assessed the clinico-pathological agreement rates. Their conclusions suggest that while histopathology remains dominant, clinical features alone may offer more value than previously assumed.
As noted above, the field of dermatology has experienced a notable increase in research aimed at improving the diagnosis and management of ESDs. As discovered by the improving predictive ability of models, diagnosing ESDs is largely considered a datamining problem.[23,24] Data mining is a complex means by which valuable information is obtained from large amounts of data. This process draws on multidisciplinary fields such as statistics, artificial intelligence, and machine learning, utilizing a variety of models and algorithms that yield diverse outcomes.[25]
More recent studies have shifted toward ensemble and hybrid machine learning models. Bozok and Çalhan [17] compared several supervised learning methods, including logistic regression, K-nearest neighbors (kNN), support vector classifier (SVC), Gaussian naïve Bayes, decision tree, and random forest. Their study found that naïve Bayes produced up to 100% classification accuracy when used with histopathological features. Wang and Xie used hybrid methods such as granular computing (GrC) and support vector machines (SVM) to draw connections between this dataset and predictive ability.[18] The increasing complexity of newer algorithms and hybrid models has fueled even more research in the field. Recently, Swain et. al. employed a hybrid ensemble model incorporating SVM, logistic regression, kNN, and decision tree classifiers.[19] Their approach involved one-way ANOVA and Chi-square tests for feature selection and GridSearchCV for hyperparameter optimization. They achieved 98.9% accuracy and further reinforced the superiority of combined feature use. Introducing a more AI-centered approach, a 2025 study conducted by Balbal [20] used a wrapper-based feature selection and compared six machine learning algorithms, again finding naïve Bayes to be the top performer with an accuracy rate of 99.45%. Most notably, this study demonstrated that even reduced feature sets involving fewer than 20 variables could still yield accuracies above 99%. This very recent research continues to affirm the ongoing interest in optimizing diagnostic tools for clinical efficiency without compromising accuracy. Complementing these modeling advances, Jogu [21] provides a broad review of current machine learning practices in ESD diagnosis. His work highlights the trend towards lightweight, explainable models that balance performance with real-world usability. These types of models help achieve the ultimate goal of providing improved quality care to patients with ESDs. Finally, Sopjani et. al.[22] contribute a clinical perspective by reviewing agreement rates between clinical and histopathological diagnoses. They reviewed 29 studies where different skin diseases were investigated and assessed the clinico-pathological agreement rates. Their conclusions suggest that while histopathology remains dominant, clinical features alone may offer more value than previously assumed. As noted above, the field of dermatology has experienced a notable increase in research aimed at improving the diagnosis and management of ESDs. As discovered by the improving predictive ability of models, diagnosing ESDs is largely considered a datamining problem.[23,24] Data mining is a complex means by which valuable information is obtained from large amounts of data. This process draws on multidisciplinary fields such as statistics, artificial intelligence, and machine learning, utilizing a variety of models and algorithms that yield diverse outcomes.[25] Despite the increasing complexity of modern analytical approaches, a consistent emphasis remains on balancing accuracy, efficiency, and clinical relevance across the UCI Dermatology dataset and similar datasets.
Statistical Analysis:
This study employed a comprehensive statistical and graphical
analysis of the Dermatology dataset sourced from the UCI Machine
Learning Repository[12] in an effort to explore and confirm existing
knowledge using R.After importing the dataset using the readxl package in R, clinical and histopathological features were grouped into two respective categories based on domain knowledge. The rowMeans() function was used to compute the average clinical and pathological scores per patient. From these averages, we generated composite predictors representing overall clinical and histopathological severity for each subject.
We performed a paired t-test using these averages to determine whether clinical and histopathological measurements differed. In addition, patients were stratified by their class label, indicating one of the six ESDs, and separate paired t-tests were conducted within each diagnostic group. The paired structure accounted for repeated measurements on the same individuals. Both histograms and boxplots indicate that the differences approximately follow a normal distribution. Our analysis of individual disease classes helped assess whether certain diseases rely more heavily on clinical or pathological characteristics for accurate classification.
Other graphical tools were implemented to supplement the statistical analyses and assess underlying assumptions. As mentioned previously, boxplots and histograms were created as summary visualizations to assess symmetry and potential deviations from normality, like skewness and outliers. Back-to-back boxplots helped compare the two groups of data and their distributions by class. We also used paired line plots as an extra method to visually assess differences within each class. These visualization techniques were not only used for exploratory purposes but also for validation of paired t-test model assumptions.
We fit six multiple logistic regression models to assess the relative predictive power of clinical and histopathological features across each ESD. For each model, the outcome variable was binary (1 = presence of a specific ESD class/disease; 0 = all other classes/diseases). The model predictors included the average clinical score and average pathological score across the respective feature sets. The models were built using the glm() function in R with a binomial family and logit link. This approach enabled the estimation of the independent contribution of clinical and pathological data in classifying disease presence, while also allowing for direct comparison of predictive trends across disease types.
All analyses were conducted using R version 4.3.1. In addition to readxl, packages utilized in this analysis include dplyr for data wrangling, ggplot2 for data visualization, tidyr for reshaping data, and ggpubr for graphical hypothesis testing annotation. These tools enabled reproducible, transparent, and efficient statistical modeling across the dataset as a whole and individual disease classes
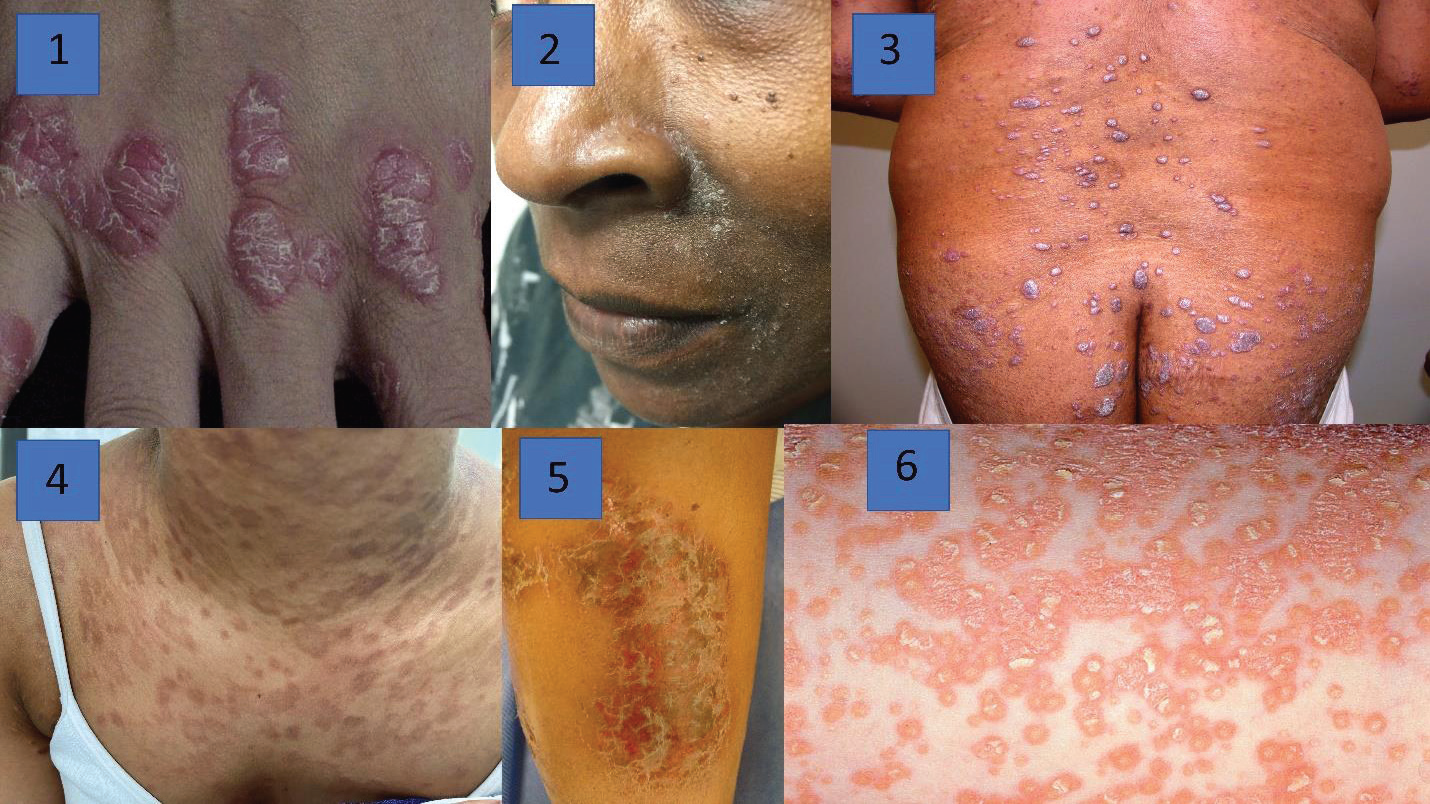
Figure 1:Clinical examples of the six ESDs studied. 1) Psoriasis; 2)
Seborrheic dermatitis; 3) Lichen planus; 4) Pityriasis rosea; PRP; 5)
Dermatitis; 6) Pityriasis rubra pilaris.
Results
Though classically the six ESDs studied differ clinically [Figure 1],
there are many times in which the diagnosis can be unclear and a skin
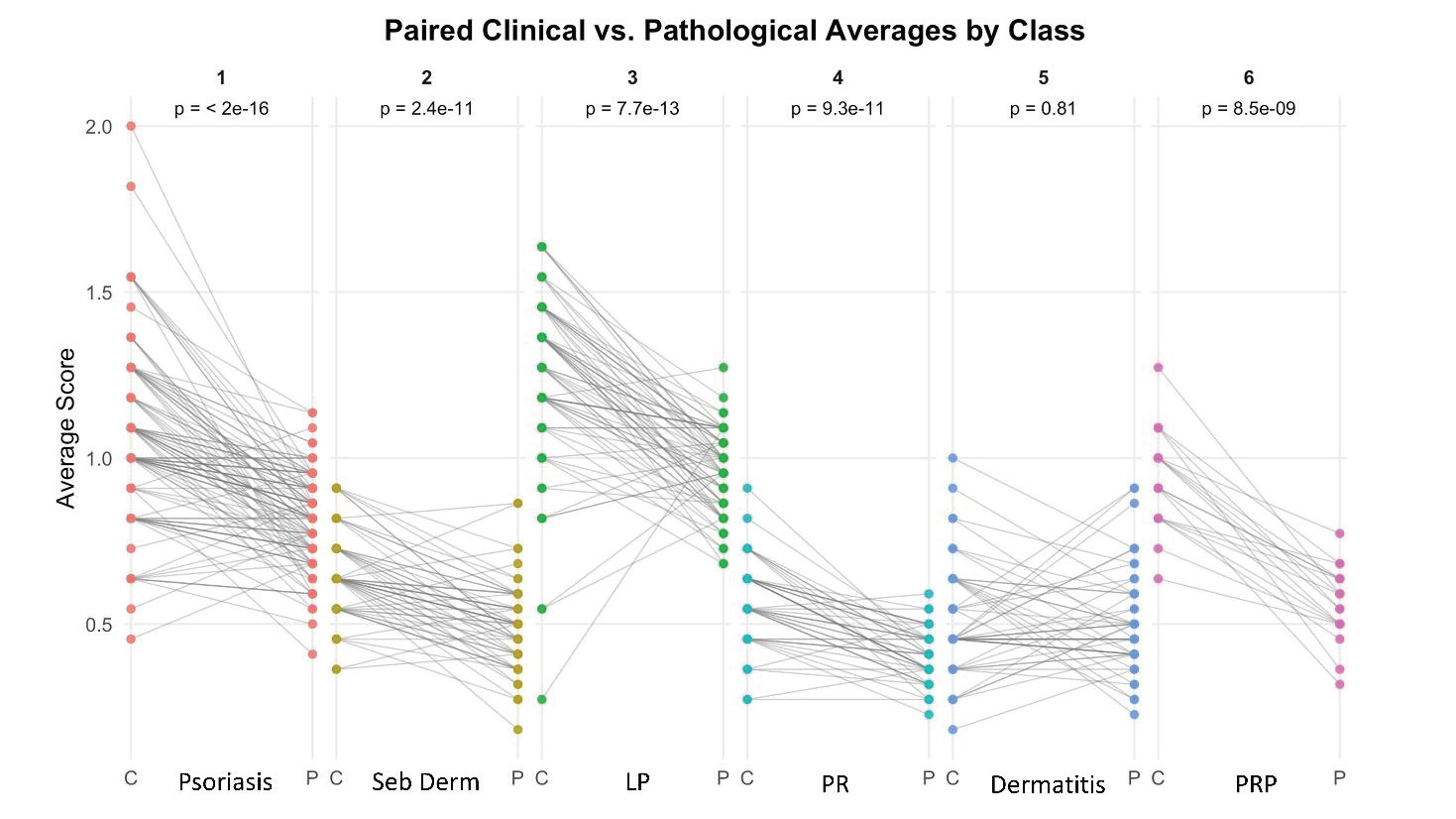
biopsy often obtained. As depicted in [Figure 2], the general paired
t-test analysis across all 366 patients revealed a statistically significant
difference between clinical and pathological averages (p < 2.2e-16),
with a mean difference of 0.202 (95% CI: 0.178–0.227). These findings
support the alternative hypothesis that the two types of features are
not equally expressed across patients. Similarly, this finding persisted
in 5 out of 6 of the individual disease classes.
- Psoriasis (Class 1): mean difference of 0.249 (p < 2.2e-16; 95% CI: 0.202–0.296)
- Seborrheic Dermatitis (Class 2): mean difference = 0.159 (p = 2.42e-11; 95% CI: 0.120–0.198)
- Lichen Planus (Class 3): mean difference of 0.286 (p = 7.71e- 13; 95% CI: 0.221–0.351)
- Pityriasis Rosea (Class 4): mean difference of 0.163 (p = 9.26e-11; 95% CI: 0.123–0.203)
- Pityriasis Rubra Pilaris (Class 6): mean difference = 0.373 (p = 8.49e-09; 95% CI: 0.292–0.453)
- Psoriasis (Class 1): mean difference of 0.249 (p < 2.2e-16; 95% CI: 0.202–0.296)
- Seborrheic Dermatitis (Class 2): mean difference = 0.159 (p = 2.42e-11; 95% CI: 0.120–0.198)
- Lichen Planus (Class 3): mean difference of 0.286 (p = 7.71e- 13; 95% CI: 0.221–0.351)
- Pityriasis Rosea (Class 4): mean difference of 0.163 (p = 9.26e-11; 95% CI: 0.123–0.203)
- Pityriasis Rubra Pilaris (Class 6): mean difference = 0.373 (p = 8.49e-09; 95% CI: 0.292–0.453)
Interestingly, the only class where no statistically significant
difference was found included Class 5 patients with Chronic
Dermatitis: mean difference = 0.007 (p = 0.810; 95% CI crossing zero
(–0.051 to 0.065)). A paired line plot was constructed to visualize
these distributions within and outside of the 6 ESD classes [Figure 2].
We fitted multiple logistic regression models to examine
the predictive capacity of averaged clinical (clin_avg) and
histopathological (hist_avg) features for each erythemato-squamous
disease (ESD) class.
For patients with psoriasis (class 1), the fitted model was:
with both predictors statistically significant (p = 0.031 for clinical
average; p < 0.001 for histopathological average). The model achieved
a predictive accuracy of 0.6202.
For patients with seborrheic dermatitis (class 2), the fitted
model was:
with only the histopathological variable significant (p = 0.0004);
the clinical average did not reach significance (p = 0.137). The model
achieved a predictive accuracy of 0.8306.
In patients with lichen planus (class 3), the fitted model was:
again, demonstrating significance for both predictors (p = 0.0008
for clinical average; p < 0.001 for histopathological average). The
model achieved a predictive accuracy of 0.8689.
For patients with pityriasis rosea (class 4), the fitted model was:
and revealed only the histopathological average to be statistically
significant (p < 0.001), whereas the clinical average approached
significance (p = 0.095). The model achieved a predictive accuracy
of 0.8661.
Interestingly, patients with chronic dermatitis (class 5) yielded
the fitted equation:
with only the clinical average reaching statistical significance (p <
0.001). The histopathological variable was not significant (p = 0.715).
The model achieved a predictive accuracy of 0.8689.
Finally, patients with pityriasis rubra pilaris (class 6)
demonstrated a strong relationship in its model:
with both predictors highly significant (p = 0.0004 for clinical
average; p < 0.001 for histopathological average). The model achieved
a predictive accuracy of 0.9454.
Discussion
The consistency and strength of our paired t-test findings suggest
that clinical manifestations tend to present more severely or distinctly
than their corresponding histopathological features. This difference
is visually illustrated in [Figure 2] and may be interpreted in several
ways. Clinically, it makes sense that patients may present with more
prominent and bothersome external symptoms in comparison to
histopathological changes, which may either lag, be more subtle, or
present overlapping patterns across diseases, reducing their apparent
distinctiveness in early or ambiguous cases. This supports the notion
that clinical features are often more immediately actionable for
diagnosis, whereas histopathology may serve better in ambiguous,
refractory, or atypical presentations.
Swain et al.[19] explored patient experience and the burden of
disease, noting that external symptoms heavily influence quality of
life. Our finding that clinical features overall appear more severe
supports the idea that visible symptoms are not only diagnostically
valuable but also impactful for patients.
Bozok and Çalhan emphasized that with histopathological
interpretation comes much subjective variability, especially for
overlapping diseases like chronic dermatitis and lichen planus.[17]
Our finding that class 5 (chronic dermatitis) lacked histopathological
significance may reflect this histopathological ambiguity.
The results of our multiple logistic regression added insight to
this paired t-test analysis. Strong positive associations were observed
in psoriasis (class 1), lichen planus (class 3), and pityriasis rubra
pilaris (class 6). In these models, both clinical and pathological
averages were statistically significant. Both predictors in these classes
had p-values < 0.05, and most < 0.001, indicating robust statistical
contributions to disease classification. These results align with prior
studies that emphasized the distinctive histopathological architecture
of these diseases, reinforcing their separability based on quantitative
feature averages.[13,20]
Uniquely, seborrheic dermatitis (class 2) showed significant
negative associations for both predictors. Only histopathological
p-values were < 0.001, suggesting that lower average values were
more indicative of this diagnosis. This may reflect the relatively milder
expression of histopathological features in seborrheic dermatitis
compared to other ESDs. Prior literature has noted the diagnostic
challenges in distinguishing seborrheic dermatitis from psoriasis,
[17] especially in early stages, which may contribute to the inverse
predictive pattern here.
In comparison, pityriasis rosea (class 4) displayed a very modest
pattern, finding the histopathological average statistically significant
(p < 0.001), while the clinical average approached significance (p =
0.095). This may suggest that histopathological features are more
diagnostic for this condition. This finding is supported by literature
noting that pityriasis rosea’s clinical features often resemble viral
exanthems, making histological confirmation particularly valuable in
ambiguous cases.[19]
Like our paired t-test results, chronic dermatitis (class 5) yielded
a unique result. It was the one class where only the clinical average
proved statistically significant. The heterogeneity and fluctuating
course of chronic dermatitis may explain this, as suggested by Elsayad
et al. (2018), who emphasized the variable inflammatory patterns
and diagnostic ambiguity of chronic dermatitis in both clinical and
microscopic contexts.[16]
The predictive accuracy of these 6 class models provided insight
into further distinguishing features of disease. Importantly, the overall
predictive performance of the models was generally high across all six
ESD classes, given the statistical simplicity of our models. It is worth
noting that some logistic regression coefficients were positive while
others were negative. However, this can be explained by the relatively
strong correlation between the clinical and histopathological features.
In every class except class 5, chronic dermatitis, both variables follow
the same direction, either positive or negative, reflecting good
correlation. Even when the coefficients differ in sign, as seen in class
5, the combined information from both predictors still contributes
meaningfully to class discrimination.
This synergistic effect likely underlies the generally high predictive accuracy observed, with particularly strong performance in classes such as pityriasis rubra pilaris (class 6), where both predictors were highly significant and closely aligned in magnitude. Overall, these results reaffirm that averaged clinical and histopathological features collectively provide substantial discriminative power for differentiating ESD classes.
Ultimately, clinical and histopathological data combined help aid in the diagnosis and treatment of these unfortunate skin diseases. The fact that histopathology alone was significant in pityriasis rosea and that the opposite held true for chronic dermatitis suggests that some diagnoses may require additional diagnostic modalities or feature engineering beyond simple averages. These insights are critical for developing machine-learning models or decision support tools that depend on interpretable, aggregated clinical data.[1,14] Further advocacy for more integrative diagnostic frameworks, combining important clinical intuition with valuable histopathology, and revolutionary AI is paramount.[21] Our data aligns well with this vision, showing that while clinical features often dominate, histopathological data remain critical for nuanced understanding, especially in ambiguous or non-classical cases.
This synergistic effect likely underlies the generally high predictive accuracy observed, with particularly strong performance in classes such as pityriasis rubra pilaris (class 6), where both predictors were highly significant and closely aligned in magnitude. Overall, these results reaffirm that averaged clinical and histopathological features collectively provide substantial discriminative power for differentiating ESD classes.
Ultimately, clinical and histopathological data combined help aid in the diagnosis and treatment of these unfortunate skin diseases. The fact that histopathology alone was significant in pityriasis rosea and that the opposite held true for chronic dermatitis suggests that some diagnoses may require additional diagnostic modalities or feature engineering beyond simple averages. These insights are critical for developing machine-learning models or decision support tools that depend on interpretable, aggregated clinical data.[1,14] Further advocacy for more integrative diagnostic frameworks, combining important clinical intuition with valuable histopathology, and revolutionary AI is paramount.[21] Our data aligns well with this vision, showing that while clinical features often dominate, histopathological data remain critical for nuanced understanding, especially in ambiguous or non-classical cases.
Conclusions
We demonstrated that averaged clinical and histopathological
features can effectively predict most erythemato-squamous disease
classes, with the strongest associations observed in lichen planus and
pityriasis rubra pilaris. While clinical severity averages emerged as
significant predictors in most disease classes, they were not universally
dominant. For conditions like lichen planus, histopathological
features provided stronger predictive power, emphasizing their
diagnostic value. This variability suggests that while clinicians may
often prioritize observable clinical features in practice, optimal
diagnosis benefits from tailored, disease-specific weighting of both
clinical and pathological data.
Exceptions, such as chronic dermatitis (class 5) prove clinical features can be more diagnostic. This underscores the importance of a good clinical exam, especially for diagnostically ambiguous or rare conditions. Our findings validate and extend prior studies using this dataset and highlight the value of hybrid, multi-modal diagnostic approaches in dermatology.
Exceptions, such as chronic dermatitis (class 5) prove clinical features can be more diagnostic. This underscores the importance of a good clinical exam, especially for diagnostically ambiguous or rare conditions. Our findings validate and extend prior studies using this dataset and highlight the value of hybrid, multi-modal diagnostic approaches in dermatology.
Author Contributions:
Conceptualization: S.S, M.G.S.; Writing-Original Draft
Preparation: M.G.S.; Writing- Review and Editing; S.S., J.B.T.;
Experimental Studies: M.G.S., S.S.; Project Administration: S.S.,
J.B.T..; Funding Acquisition: J.B.T.Acknowledgements:
This research was supported in part by grants from the National
Institutes of Health (R01 HL062996 (J.B.T.), R01 ES031087 (J.B.T.),
and an unrestricted educational grant from the Wright State
University Department of Pharmacology & Toxicology.Conflicts of Interest:
The authors declare no conflicts of interest.References
Citation
Shenouda MG, Travers JB, Sun S. Comparing Clinical vs Histopathological Features in Diagnosing Erythemato-Squamous Diseases. J Clin Investigat Dermatol. 2026;14(1): 1