Advances in Diabetes & Endocrinology
Download PDF
Among these, DNA methylation is the most extensively studied and clinically validated mechanism, involving the addition of methyl groups to cytosine residues at CpG dinucleotides, typically leading to transcriptional repression. Histone modifications, including acetylation, methylation, phosphorylation, and ubiquitination, influence chromatin structure and accessibility, thereby regulating gene transcription. Non-coding RNAs further contribute to posttranscriptional and transcriptional gene regulation by modulating mRNA stability and protein translation.
In contrast to fixed genetic mutations, epigenetic modifications are dynamic and responsive to environmental influences such as diet, physical activity, obesity, and pharmacological interventions. This plasticity allows the epigenome to reflect cumulative metabolic exposures over time. Importantly, these changes can be detected in accessible biological samples, including peripheral blood and saliva, facilitating their application in clinical settings.
Collectively, these characteristics position epigenetic mechanisms as highly relevant to precision medicine. Their reversibility and sensitivity to therapeutic and lifestyle interventions offer promising opportunities not only for early disease detection and risk stratification but also for the development of targeted strategies aimed at disease modification and potential reversal.
Furthermore, lipid accumulation induces lipotoxicity, promoting oxidative stress, mitochondrial dysfunction, and activation of inflammatory signalling pathways. These processes impair β-cell viability and function by reducing insulin synthesis and secretion. Over time, sustained metabolic stress contributes to β-cell apoptosis and progressive decline in insulin-producing capacity. In addition to its direct effects on β-cells, altered ABCG1 methylation in peripheral tissues such as adipose tissue and macrophages exacerbates systemic insulin resistance through dysregulated lipid metabolism and chronic low-grade inflammation. This creates a dual pathogenic effect—both impaired insulin secretion and increased insulin resistance—central to the development of T2DM.
Clinically, the methylation status of ABCG1 (cg06500161) in peripheral blood has shown strong and reproducible associations with incident T2DM across multiple populations. Its role as an early biomarker reflects underlying metabolic dysregulation before overt hyperglycemia, making it a valuable tool for early risk prediction and intervention.
In β-cells, elevated TXNIP expression contributes to impaired insulin secretion through multiple mechanisms. These include mitochondrial dysfunction, activation of apoptotic pathways, and suppression of insulin gene transcription. Additionally, TXNIP has been shown to activate the NLRP3 inflammasome, leading to the production of pro-inflammatory cytokines such as interleukin-1β (IL-1β), further exacerbating β-cell damage and dysfunction.
Beyond the pancreas, TXNIP plays a role in peripheral insulin resistance. Increased TXNIP expression in skeletal muscle and adipose tissue interferes with glucose uptake and insulin signalling pathways, thereby contributing to systemic hyperglycemia. Importantly, TXNIP expression is directly induced by high glucose levels, creating a feedforward pathogenic loop: hyperglycemia increases TXNIP expression, which in turn worsens oxidative stress and β-cell dysfunction, leading to further elevations in blood glucose levels.
Clinically, hypomethylation of TXNIP (cg19693031) in peripheral blood has been consistently associated with elevated fasting glucose and HbA1c levels, making it a robust biomarker of poor glycemic control. Its dynamic nature also allows it to reflect ongoing metabolic status and response to therapeutic interventions.
• PPARGC1A – Regulates mitochondrial biogenesis and oxidative metabolism, playing a central role in maintaining insulin sensitivity in skeletal muscle. Hypermethylation of PPARGC1A is associated with reduced gene expression, leading to impaired mitochondrial function and contributing to insulin resistance in T2DM.
• PDX-1 – A key transcription factor essential for pancreatic β-cell development and insulin gene expression. Epigenetic silencing through hypermethylation reduces PDX-1 activity, resulting in impaired insulin secretion and progressive β-cell dysfunction.
• PDGFA – Involved in cell growth and tissue remodelling, particularly in hepatic metabolism. Altered methylation of PDGFA is associated with increased liver fat accumulation and hepatic insulin resistance, contributing to metabolic dysregulation in T2DM.Thousands of differentially methylated loci have been identified across adipose tissue, skeletal muscle, and liver, reflecting systemic metabolic dysfunction.
A major advancement in epigenetic research is the ability to predict long-term complications of T2DM, particularly cardiovascular disease and diabetic kidney disease. Epigenetic alterations reflect cumulative metabolic stress and may precede clinical manifestations, enabling early identification of high-risk individuals. Epigenetic modifications in genes regulating inflammation, lipid metabolism, and endothelial function have been strongly associated with increased cardiovascular risk in T2DM. Large-scale epigenomewide association studies (EWAS) have identified differential DNA methylation patterns linked to incident cardiovascular events. For example, a study by García-Calzón et al. (2025, Cell Reports Medicine) demonstrated that specific methylation signatures in circulating DNA could predict major adverse cardiovascular events (MACE), including myocardial infarction and ischemic stroke, in patients with T2DM. These findings suggest that epigenetic profiling can stratify patients beyond traditional risk factors such as lipid levels and blood pressure.
• miR-375 – A key regulator of pancreatic β-cell function, miR-375 modulates insulin secretion and β-cell mass.
Dysregulation contributes to impaired insulin release and progression of T2DM.
• miR-29 family – These miRNAs are associated with insulin resistance by targeting genes involved in glucose uptake and insulin signalling pathways, particularly in skeletal muscle and adipose tissue.
• miR-126 – Primarily expressed in endothelial cells, miR-126 plays a crucial role in vascular integrity and angiogenesis. Reduced levels are linked to endothelial dysfunction and increased risk of diabetic vascular complications.
• MALAT1 – A regulator of inflammation and endothelial function, MALAT1 is upregulated in hyperglycemic states and contributes to vascular damage and diabetic complications. These circulating non-coding RNAs are highly stable in body fluids such as blood and plasma, making them suitable for noninvasive “liquid biopsy” approaches for early diagnosis, prognosis, and monitoring of T2DM.
Epigenetic markers significantly improve 10-year risk prediction models, enabling:
• Identification of high-risk individuals
• Prevention of progression from prediabetes to T2DM
Complication Prediction
Distinct epigenetic profiles help classify T2DM into subgroups such as:
• Severe insulin-resistant diabetes
• High cardiovascular risk phenotypes
This allows prediction of:
• Heart attack
• Stroke
• Kidney disease
Monitoring Disease Evolution
Dynamic changes in DNA methylation track:
• Transition from normoglycemia → prediabetes → diabetes
• Progression of metabolic dysfunction
• Lifestyle interventions (diet, exercise)
• Pharmacological treatment (e.g., metformin)
• Real-Time Metabolic Insight: Unlike HbA1c, which reflects average glycemic control over the preceding 2–3 months, epigenetic markers provide insights into ongoing metabolic alterations and tissue-specific dysfunction, enabling a more dynamic assessment of disease activity.
• Metabolic Memory: Epigenetic modifications capture the long-term effects of prior hyperglycemic exposure, reflecting the concept of “metabolic memory,” whereby earlier glycemic insults continue to influence disease progression and complications.
• Precision Medicine: Epigenetic profiling allows stratification of patients into distinct molecular subtypes, facilitating individualized therapeutic approaches and improving clinical decision-making.
• Non-Invasive Testing: Advances in liquid biopsy techniques enable the detection of circulating cell-free DNA (cfDNA) methylation and other epigenetic markers in blood and saliva, offering a minimally invasive and clinically feasible diagnostic approach.
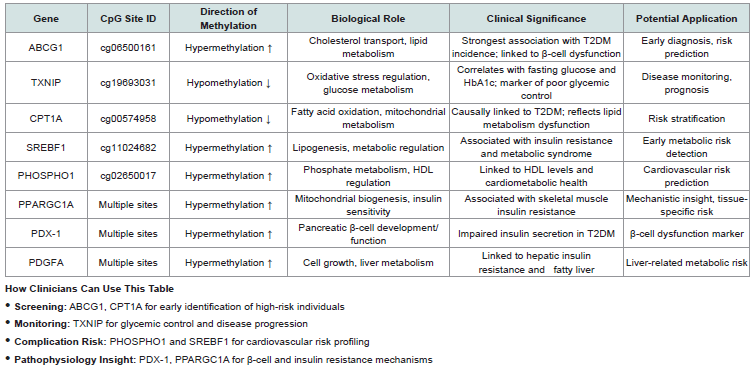
[Table 1] summarizes the key CpG DNA methylation biomarkers in type 2 diabetes mellitus and highlights their potential clinical applications in diagnosis, risk assessment, and disease monitoring.
• Diet – alters methylation of metabolic genes
• Obesity – induces widespread epigenetic dysregulation
• Physical activity – can reverse adverse epigenetic patterns
• Intrauterine exposure – influences lifelong metabolic risk
This reinforces the importance of early lifestyle interventions in clinical practice.
Review Article
Epigenetic Biomarkers in Type 2 Diabetes: A Clinician’s Guide to Emerging Diagnostic and Prognostic Tools
Sibi Das*
Department of Medicine, NC Medical College, Israna, Panipat, Haryana, India.
*Address for Correspondence:Dr. Sibi Das, Department of Medicine, NC Medical College,
Israna, Panipat, Haryana, India. E-mail Id: sdsilvanose@gmail.com
Submission:27 March, 2026
Accepted:21 April, 2026
Published:23 April, 2026
Copyright: © 2026 Das S This is an open access article
distributed under the Creative Commons Attribution License,
which permits unrestricted use, distribution, and reproduction
in any medium, provided the original work is properly cited.
Keywords:Type 2 diabetes mellitus; Epigenetics; DNA methylation; CpG sites;
Biomarkers; Precision medicine; Diabetic complications; Cardiovascular risk
Abstract
Type 2 diabetes mellitus (T2DM) is a chronic metabolic disorder characterized
by insulin resistance and progressive pancreatic β-cell dysfunction. Conventional
biomarkers such as fasting plasma glucose and glycated haemoglobin (HbA1c)
detect established disease but fail to capture early molecular alterations and
future complication risk. Epigenetic biomarkers—including DNA methylation, histone
modifications, and non-coding RNAs—are emerging as powerful, dynamic, and
modifiable indicators of disease susceptibility and progression. These biomarkers not
only enable early diagnosis and risk stratification but also hold promise in predicting
major complications such as myocardial infarction, stroke, and diabetic kidney disease.
This clinician-focused review summarizes current evidence on epigenetic biomarkers
in T2DM, highlights key CpG sites, and discusses their diagnostic, prognostic, and
therapeutic implications in precision medicine.
Introduction
Type 2 diabetes mellitus (T2DM) represents a major and rapidly
escalating global health burden. According to the International
Diabetes Federation (IDF), approximately 537 million adults
worldwide were living with diabetes in 2021, a number projected to
rise to 643 million by 2030 and 783 million by 2045. Notably, over
90-95% of these cases are attributed to T2DM, underscoring its
dominant contribution to the global diabetes epidemic. The disease
is associated with substantial morbidity and mortality, primarily due
to complications such as cardiovascular disease, stroke, and chronic
kidney disease, which remain the leading causes of death among
diabetic patients.
In addition to its clinical burden, T2DM imposes significant economic strain, with global health expenditure on diabetes estimated to exceed USD 966 billion annually, reflecting both direct healthcare costs and indirect productivity losses. Alarmingly, a large proportion of individuals remain undiagnosed or are diagnosed late, often after the onset of irreversible vascular and metabolic complications. Current diagnostic tools, including fasting plasma glucose and glycated haemoglobin (HbA1c), primarily detect hyperglycemia after substantial metabolic dysfunction has already occurred. These conventional markers are limited in their ability to identify early molecular changes, predict disease progression, or stratify risk for complications.
Emerging evidence suggests that epigenetic mechanisms act as a critical interface between genetic predisposition and environmental influences such as diet, obesity, physical inactivity, and intrauterine exposures. These modifications—including DNA methylation, histone alterations, and non-coding RNA regulation—can precede clinical disease onset and reflect cumulative metabolic stress. Importantly, epigenetic changes are dynamic and potentially reversible, offering a unique opportunity for early detection, risk stratification, and targeted intervention. From a clinical perspective, epigenetic biomarkers may enable the identification of high-risk individuals before the development of overt hyperglycemia, predict the likelihood of complications, and guide personalized therapeutic strategies. As such, they represent a promising frontier in the transition toward precision medicine in T2DM.
In addition to its clinical burden, T2DM imposes significant economic strain, with global health expenditure on diabetes estimated to exceed USD 966 billion annually, reflecting both direct healthcare costs and indirect productivity losses. Alarmingly, a large proportion of individuals remain undiagnosed or are diagnosed late, often after the onset of irreversible vascular and metabolic complications. Current diagnostic tools, including fasting plasma glucose and glycated haemoglobin (HbA1c), primarily detect hyperglycemia after substantial metabolic dysfunction has already occurred. These conventional markers are limited in their ability to identify early molecular changes, predict disease progression, or stratify risk for complications.
Emerging evidence suggests that epigenetic mechanisms act as a critical interface between genetic predisposition and environmental influences such as diet, obesity, physical inactivity, and intrauterine exposures. These modifications—including DNA methylation, histone alterations, and non-coding RNA regulation—can precede clinical disease onset and reflect cumulative metabolic stress. Importantly, epigenetic changes are dynamic and potentially reversible, offering a unique opportunity for early detection, risk stratification, and targeted intervention. From a clinical perspective, epigenetic biomarkers may enable the identification of high-risk individuals before the development of overt hyperglycemia, predict the likelihood of complications, and guide personalized therapeutic strategies. As such, they represent a promising frontier in the transition toward precision medicine in T2DM.
Epigenetics: A Clinician-Oriented Overview:
Epigenetics refers to heritable yet reversible modifications in
gene expression that occur without alterations in the underlying
DNA sequence. These regulatory mechanisms play a crucial
role in modulating cellular function and phenotype in response
to environmental and metabolic cues. The principal epigenetic
mechanisms include DNA methylation, histone modifications, and
regulation by non-coding RNAs such as microRNAs (miRNAs) and
long non-coding RNAs (lncRNAs).Among these, DNA methylation is the most extensively studied and clinically validated mechanism, involving the addition of methyl groups to cytosine residues at CpG dinucleotides, typically leading to transcriptional repression. Histone modifications, including acetylation, methylation, phosphorylation, and ubiquitination, influence chromatin structure and accessibility, thereby regulating gene transcription. Non-coding RNAs further contribute to posttranscriptional and transcriptional gene regulation by modulating mRNA stability and protein translation.
In contrast to fixed genetic mutations, epigenetic modifications are dynamic and responsive to environmental influences such as diet, physical activity, obesity, and pharmacological interventions. This plasticity allows the epigenome to reflect cumulative metabolic exposures over time. Importantly, these changes can be detected in accessible biological samples, including peripheral blood and saliva, facilitating their application in clinical settings.
Collectively, these characteristics position epigenetic mechanisms as highly relevant to precision medicine. Their reversibility and sensitivity to therapeutic and lifestyle interventions offer promising opportunities not only for early disease detection and risk stratification but also for the development of targeted strategies aimed at disease modification and potential reversal.
DNA Methylation: Core Epigenetic Biomarkers in T2DM:
DNA methylation at specific CpG (cytosine-guanine
dinucleotides) sites has emerged as the most robust and clinically
relevant epigenetic biomarker in T2DM.Key CpG Sites and Their Clinical Significance:
Research has identified several CpG loci detectable in blood long
before clinical disease onset:• ABCG1 (cg06500161):
Strongest association with T2DM incidence; hypermethylation
linked to impaired insulin secretion and β-cell dysfunction. ABCG1
(ATP-binding cassette transporter G1) plays a critical role in
intracellular cholesterol homeostasis by facilitating cholesterol efflux
to high-density lipoproteins (HDL). Proper cholesterol balance is
essential for maintaining cellular membrane integrity and normal
metabolic signaling, particularly in pancreatic β-cells.
Hypermethylation at the CpG site cg06500161 within the
ABCG1 gene is associated with reduced gene expression. This
epigenetic silencing leads to impaired cholesterol efflux, resulting in
intracellular lipid and cholesterol accumulation. In pancreatic β-cells,
excess cholesterol disrupts membrane microdomains, including lipid
rafts, which are crucial for insulin granule exocytosis and glucose stimulated
insulin secretion.Furthermore, lipid accumulation induces lipotoxicity, promoting oxidative stress, mitochondrial dysfunction, and activation of inflammatory signalling pathways. These processes impair β-cell viability and function by reducing insulin synthesis and secretion. Over time, sustained metabolic stress contributes to β-cell apoptosis and progressive decline in insulin-producing capacity. In addition to its direct effects on β-cells, altered ABCG1 methylation in peripheral tissues such as adipose tissue and macrophages exacerbates systemic insulin resistance through dysregulated lipid metabolism and chronic low-grade inflammation. This creates a dual pathogenic effect—both impaired insulin secretion and increased insulin resistance—central to the development of T2DM.
Clinically, the methylation status of ABCG1 (cg06500161) in peripheral blood has shown strong and reproducible associations with incident T2DM across multiple populations. Its role as an early biomarker reflects underlying metabolic dysregulation before overt hyperglycemia, making it a valuable tool for early risk prediction and intervention.
• TXNIP (cg19693031):
Hypomethylation correlates with elevated fasting glucose and
HbA1c, reflecting poor glycemic control. TXNIP (Thioredoxin-
Interacting Protein) is a key regulator of cellular redox balance and
glucose metabolism. It binds to and inhibits thioredoxin, a major
intracellular antioxidant, thereby promoting oxidative stress.
Hypomethylation at the CpG site cg19693031 within the TXNIP
gene is associated with increased gene expression. This epigenetic
upregulation leads to elevated TXNIP protein levels, which in turn
inhibit thioredoxin activity and result in the accumulation of reactive
oxygen species (ROS). Increased oxidative stress has detrimental
effects on pancreatic β-cells, which are particularly vulnerable due to
their relatively low antioxidant capacity.In β-cells, elevated TXNIP expression contributes to impaired insulin secretion through multiple mechanisms. These include mitochondrial dysfunction, activation of apoptotic pathways, and suppression of insulin gene transcription. Additionally, TXNIP has been shown to activate the NLRP3 inflammasome, leading to the production of pro-inflammatory cytokines such as interleukin-1β (IL-1β), further exacerbating β-cell damage and dysfunction.
Beyond the pancreas, TXNIP plays a role in peripheral insulin resistance. Increased TXNIP expression in skeletal muscle and adipose tissue interferes with glucose uptake and insulin signalling pathways, thereby contributing to systemic hyperglycemia. Importantly, TXNIP expression is directly induced by high glucose levels, creating a feedforward pathogenic loop: hyperglycemia increases TXNIP expression, which in turn worsens oxidative stress and β-cell dysfunction, leading to further elevations in blood glucose levels.
Clinically, hypomethylation of TXNIP (cg19693031) in peripheral blood has been consistently associated with elevated fasting glucose and HbA1c levels, making it a robust biomarker of poor glycemic control. Its dynamic nature also allows it to reflect ongoing metabolic status and response to therapeutic interventions.
• CPT1A (cg00574958):
CPT1A (Carnitine Palmitoyltransferase 1A) is a key
mitochondrial enzyme regulating the transport of long-chain fatty
acids into mitochondria for β-oxidation. Hypomethylation at
cg00574958 is associated with increased gene expression, reflecting
altered fatty acid metabolism. Dysregulation of CPT1A activity leads
to impaired lipid oxidation and accumulation of intracellular lipids,
contributing to insulin resistance in liver and skeletal muscle. This
metabolic imbalance promotes hepatic steatosis and disrupts glucose
homeostasis, establishing a causal link with T2DM development.
Clinically, CPT1A methylation serves as an indicator of early lipid
metabolic dysfunction and cardiometabolic risk.• SREBF1 (cg11024682):
SREBF1 (Sterol Regulatory Element-Binding Transcription
Factor 1) is a central regulator of lipogenesis, controlling the
expression of genes involved in fatty acid and triglyceride synthesis.
Hypermethylation at cg11024682 influences gene expression
and is associated with dysregulated lipid metabolism. Altered
SREBF1 activity contributes to increased hepatic lipid synthesis,
insulin resistance, and features of metabolic syndrome, including
dyslipidemia and central obesity. These changes exacerbate systemic
metabolic stress and promote progression toward T2DM. Clinically,
SREBF1 methylation is a marker of lipogenic activity and metabolic
syndrome risk.• PHOSPHO1(cg02650017):
PHOSPHO1 (Phosphatase, Orphan 1) is involved in phosphate
metabolism and has emerging roles in energy balance and lipid
metabolism. Hypermethylation at cg02650017 has been associated
with variations in high-density lipoprotein (HDL) cholesterol levels
and overall metabolic health. Although its exact role in T2DM
pathogenesis is still being elucidated, PHOSPHO1 methylation
appears to reflect alterations in lipid handling and cardiovascular risk
profiles. Clinically, it may serve as a biomarker for cardiometabolic
health and HDL-related protective pathways.Tissue-Specific Epigenetic Markers:
Although blood-based markers are clinically convenient, tissue level
studies provide mechanistic insights:• PPARGC1A – Regulates mitochondrial biogenesis and oxidative metabolism, playing a central role in maintaining insulin sensitivity in skeletal muscle. Hypermethylation of PPARGC1A is associated with reduced gene expression, leading to impaired mitochondrial function and contributing to insulin resistance in T2DM.
• PDX-1 – A key transcription factor essential for pancreatic β-cell development and insulin gene expression. Epigenetic silencing through hypermethylation reduces PDX-1 activity, resulting in impaired insulin secretion and progressive β-cell dysfunction.
• PDGFA – Involved in cell growth and tissue remodelling, particularly in hepatic metabolism. Altered methylation of PDGFA is associated with increased liver fat accumulation and hepatic insulin resistance, contributing to metabolic dysregulation in T2DM.Thousands of differentially methylated loci have been identified across adipose tissue, skeletal muscle, and liver, reflecting systemic metabolic dysfunction.
Epigenetic Biomarkers for Prediction of Diabetic Complications:
Cardiovascular Risk (Heart Attack and Stroke)A major advancement in epigenetic research is the ability to predict long-term complications of T2DM, particularly cardiovascular disease and diabetic kidney disease. Epigenetic alterations reflect cumulative metabolic stress and may precede clinical manifestations, enabling early identification of high-risk individuals. Epigenetic modifications in genes regulating inflammation, lipid metabolism, and endothelial function have been strongly associated with increased cardiovascular risk in T2DM. Large-scale epigenomewide association studies (EWAS) have identified differential DNA methylation patterns linked to incident cardiovascular events. For example, a study by García-Calzón et al. (2025, Cell Reports Medicine) demonstrated that specific methylation signatures in circulating DNA could predict major adverse cardiovascular events (MACE), including myocardial infarction and ischemic stroke, in patients with T2DM. These findings suggest that epigenetic profiling can stratify patients beyond traditional risk factors such as lipid levels and blood pressure.
Similarly, methylation changes in genes such as ABCG1, SREBF1,
and inflammatory pathway regulators have been associated with
atherosclerosis progression and endothelial dysfunction. Altered
methylation in these pathways contributes to chronic vascular
inflammation, plaque instability, and thrombosis.
In addition, studies published in high-impact cardiovascular
and metabolic journals have shown that epigenetic markers related
to oxidative stress and immune activation correlate with subclinical
atherosclerosis, measured by carotid intima-media thickness and
coronary artery calcification.
Importantly, these epigenetic signatures can be detected in
peripheral blood, allowing for non-invasive early identification of
individuals at high risk of myocardial infarction and stroke, even
before overt cardiovascular disease develops.
Diabetic Kidney Disease:
Diabetic kidney disease (DKD) is one of the most serious
microvascular complications of T2DM and a leading cause of endstage
renal disease worldwide. Epigenetic mechanisms, particularly
DNA methylation, play a key role in its pathogenesis. A landmark
study by Li et al. (2023, Nature Communications) identified specific
DNA methylation markers associated with decline in renal function
and progression of DKD. These methylation signatures were shown
to predict changes in estimated glomerular filtration rate (eGFR) and
albuminuria, independent of traditional clinical risk factors.
Epigenetic alterations in genes involved in fibrosis (e.g., TGF-β
signalling), inflammation, and extracellular matrix remodelling have
been consistently linked to renal injury. These changes contribute
to Glomerulosclerosis, Tubulointerstitial fibrosis and Progressive
loss of renal function. Furthermore, studies have demonstrated that
epigenetic “metabolic memory” plays a role in DKD progression,
where prior hyperglycemic exposure induces persistent epigenetic
changes that continue to drive renal damage even after glycemic
control improves. These findings highlight the potential of DNA
methylation markers as early predictors of renal dysfunction and
progression to chronic kidney disease, enabling earlier intervention.Clinical Implication:
The integration of epigenetic biomarkers into clinical practice
offers significant advantages in predicting and preventing diabetic
complications. An epigenetic biomarker capable of identifying
complication risk at an early stage allows for:
• Preventive interventions, including early initiation of
cardioprotective and renoprotective therapies.
• Aggressive risk factor modification, such as tighter glycemic,
lipid, and blood pressure control.
• Targeted monitoring strategies, including more frequent
cardiovascular and renal assessments.
Moreover, epigenetic profiling may help classify patients into
distinct risk phenotypes, enabling a precision medicine approach
where treatment strategies are tailored according to individual risk
profiles. As evidence continues to grow, epigenetic biomarkers have
the potential to complement existing clinical tools and transform the
management of T2DM by shifting the focus from reactive treatment
to proactive prevention of complications.Non-Coding RNAs as Circulating Biomarkers:
MicroRNAs (miRNAs)• miR-375 – A key regulator of pancreatic β-cell function, miR-375 modulates insulin secretion and β-cell mass.
Dysregulation contributes to impaired insulin release and progression of T2DM.
• miR-29 family – These miRNAs are associated with insulin resistance by targeting genes involved in glucose uptake and insulin signalling pathways, particularly in skeletal muscle and adipose tissue.
• miR-126 – Primarily expressed in endothelial cells, miR-126 plays a crucial role in vascular integrity and angiogenesis. Reduced levels are linked to endothelial dysfunction and increased risk of diabetic vascular complications.
Long Non-Coding RNAs (lncRNAs):
• H19 – Involved in glucose metabolism and insulin
signalling, H19 regulates gene expression through epigenetic
mechanisms and is implicated in metabolic homeostasis and
insulin sensitivity.• MALAT1 – A regulator of inflammation and endothelial function, MALAT1 is upregulated in hyperglycemic states and contributes to vascular damage and diabetic complications. These circulating non-coding RNAs are highly stable in body fluids such as blood and plasma, making them suitable for noninvasive “liquid biopsy” approaches for early diagnosis, prognosis, and monitoring of T2DM.
Clinical Applications of Epigenetic Biomarkers:
Early Risk AssessmentEpigenetic markers significantly improve 10-year risk prediction models, enabling:
• Identification of high-risk individuals
• Prevention of progression from prediabetes to T2DM
Complication Prediction
Distinct epigenetic profiles help classify T2DM into subgroups such as:
• Severe insulin-resistant diabetes
• High cardiovascular risk phenotypes
This allows prediction of:
• Heart attack
• Stroke
• Kidney disease
Monitoring Disease Evolution
Dynamic changes in DNA methylation track:
• Transition from normoglycemia → prediabetes → diabetes
• Progression of metabolic dysfunction
Evaluation of Therapeutic Response:
Because epigenetic modifications are reversible, they can monitor:• Lifestyle interventions (diet, exercise)
• Pharmacological treatment (e.g., metformin)
Advantages Over Traditional Biomarkers:
Epigenetic biomarkers offer several advantages over conventional
markers such as glycated hemoglobin (HbA1c) and fasting
plasma glucose, particularly in the context of early detection, risk
stratification, and personalized management of T2DM.• Real-Time Metabolic Insight: Unlike HbA1c, which reflects average glycemic control over the preceding 2–3 months, epigenetic markers provide insights into ongoing metabolic alterations and tissue-specific dysfunction, enabling a more dynamic assessment of disease activity.
• Metabolic Memory: Epigenetic modifications capture the long-term effects of prior hyperglycemic exposure, reflecting the concept of “metabolic memory,” whereby earlier glycemic insults continue to influence disease progression and complications.
• Precision Medicine: Epigenetic profiling allows stratification of patients into distinct molecular subtypes, facilitating individualized therapeutic approaches and improving clinical decision-making.
• Non-Invasive Testing: Advances in liquid biopsy techniques enable the detection of circulating cell-free DNA (cfDNA) methylation and other epigenetic markers in blood and saliva, offering a minimally invasive and clinically feasible diagnostic approach.
[Table 1] summarizes the key CpG DNA methylation biomarkers in type 2 diabetes mellitus and highlights their potential clinical applications in diagnosis, risk assessment, and disease monitoring.
Environmental and Lifestyle Influence:
Epigenetic biomarkers highlight the impact of environmental
exposures:• Diet – alters methylation of metabolic genes
• Obesity – induces widespread epigenetic dysregulation
• Physical activity – can reverse adverse epigenetic patterns
• Intrauterine exposure – influences lifelong metabolic risk
This reinforces the importance of early lifestyle interventions in clinical practice.
Clinical Limitations and Challenges:
Despite promising advances, several barriers remain:Tissue Specificity:
Epigenetic changes vary across tissues, making it difficultto directly correlate blood findings with pancreatic or hepatic
dysfunction.
Causality vs Consequence
It remains unclear whether epigenetic changes:
• Cause T2DM
• Or result from hyperglycemia
Need for Validation
• Most studies are small or retrospective
• Large-scale, longitudinal studies are required
• Standardization of assays and thresholds is lacking
Future Directions
• Development of standardized epigenetic panels for clinical use
• Integration with genomics, proteomics, and clinical data
• Application of artificial intelligence for risk prediction
• Exploration of epigenetic therapies targeting reversible modifications
It remains unclear whether epigenetic changes:
• Cause T2DM
• Or result from hyperglycemia
Need for Validation
• Most studies are small or retrospective
• Large-scale, longitudinal studies are required
• Standardization of assays and thresholds is lacking
Future Directions
• Development of standardized epigenetic panels for clinical use
• Integration with genomics, proteomics, and clinical data
• Application of artificial intelligence for risk prediction
• Exploration of epigenetic therapies targeting reversible modifications
Conclusion
Epigenetic biomarkers, particularly DNA methylation, are
emerging as powerful tools in the early diagnosis, risk stratification,
and prognosis of T2DM and its complications. Their dynamic and
modifiable nature offers a unique opportunity for preventive and
personalized medicine. Importantly, the ability to predict major
complications such as cardiovascular disease and kidney dysfunction
at an early stage could transform clinical management. However,
further validation and standardization are essential before routine
clinical implementation.
References
Citation
Das S. Epigenetic Biomarkers in Type 2 Diabetes: A Clinician’s Guide to Emerging Diagnostic and Prognostic Tools. Adv Diabetes Endocrinol 2026;9(1): 1