Journal of Toxins
Download PDF
Short Communication
*Address for Correspondence: Dhananjaya BL, Toxinology/Toxicology and Drug Discovery Unit, Center for Emerging Technologies, Jain Global Campus, Jain University, Kanakapura-562112, Karnataka, India, Tel: 08197324276; E-mail: chandu_greeshma@rediffmail.com
Citation: Saikumari YK, D`Souza CJM, Dhananjaya BL. Geographic Variation in the Peptidome Fraction of the Venom of Naja naja naja (Indian Cobra) Species as Analysed by MALDI-TOF; Implications on Antivenin Development. J Toxins. 2015;2(2): 4.
Copyright © 2015 Saikumari YK, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Journal of Toxins | ISSN: 2328-1723 | Volume: 2, Issue: 2
Submission: 18 August 2015 | Accepted: 08 September 2015 | Published: 14 September 2015
Geographic Variation in the Peptidome Fraction of the Venom of Naja naja naja (Indian Cobra) Species as Analysed by MALDITOF; Implications on Antivenin Development
Saikumari YK1,2, Cletus JM D`Souza1 and Dhananjaya BL1-3*
- 1Department of Studies in Biochemistry, University of Mysore, Mysore-570006, India
- 2Molecular Biophysics Unit, Indian Institute of Science, Bangalore-560 012, India
- 3Toxinology/Toxicology and Drug Discovery Unit, Centre for Emerging Technologies, Jain Global Campus, Jain University, Kanakapura-562112, Karnataka, India
*Address for Correspondence: Dhananjaya BL, Toxinology/Toxicology and Drug Discovery Unit, Center for Emerging Technologies, Jain Global Campus, Jain University, Kanakapura-562112, Karnataka, India, Tel: 08197324276; E-mail: chandu_greeshma@rediffmail.com
Citation: Saikumari YK, D`Souza CJM, Dhananjaya BL. Geographic Variation in the Peptidome Fraction of the Venom of Naja naja naja (Indian Cobra) Species as Analysed by MALDI-TOF; Implications on Antivenin Development. J Toxins. 2015;2(2): 4.
Copyright © 2015 Saikumari YK, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Journal of Toxins | ISSN: 2328-1723 | Volume: 2, Issue: 2
Submission: 18 August 2015 | Accepted: 08 September 2015 | Published: 14 September 2015
Abstract
Purpose: Several studies have shown that the commercially available antivenoms are ineffective in neutralizing the toxic and lethal effects. In this study, we aimed at analysing the comparative peptidome fractions of venoms of Indian Cobra (Naja naja) species from three distinct geographical (western, southern and eastern) egions of India. MALDI spectra of the regional venoms were recorded in positive ion mode using a Bruker Daltonics Ultraflex TOF/TOF spectrometer.Results: It was observed that the peptidome fraction of Naja naja species of different regions, varied greatly in MALDI-TOF spectral profile in the 1-40 KDa window. The peptide pattern of the three regional venoms had similarities in containing 27, 20, 13 and 6.7 KDa toxin components. Further, it was interesting to note that the eastern regional venom sample contained an abundant ~5.7 KDa peptide which was completely absent in other regional venoms, which might be the reason for high toxic and lethal effect.
Conclusion: The geographical venom variability in peptidome fraction reported here may have an impact in the selection of specimens for antivenom production and highlights the necessity of use of pooled venoms as representative venom for antivenom production.
Keywords
Mass spectrometry; Proteome; Snake venom; Antivenom; Regional variation; 3FTxsAbbreviations
MALDI-TOF: Matrix-assisted Laser Desorption/Ionization- Time of Flight; KDa: Kilodalton; %: Percentage; μg: Microgram; μl: Microlitre; μM: MicromolarIntroduction
Snake venom constitutes a diverse and synergistic cocktail of biologically active molecules responsible for various pharmacological effects [1,2]. In this respect, venoms of snakes are known to exhibit marked variation in their potency and the extent of induction of toxic and lethal effects due to variation of toxins have been addressed at different levels (sex, diet, seasonal, geographical etc.,), and also in terms of their composition and relative abundance of toxins [3,4]. The variation in venom composition is known to be one of the main reasons for inefficiency of antivenoms, which is the only preferred choice for snakebite victim’s treatment all over the world [5]. It has been observed that the variable composition of venom influences the effectiveness of antivenom as antivenom prepared against particular regional venom is reported to be ineffective or partially effective against the toxicity/lethality of other regional venom [6-8]. Hence, understanding the intra-specific variability of venom components are gaining much attention with the intention of production of efficacious therapeutic antivenom and thus help in management of snakebite [8]. Although variation in the venom proteome is a well-documented phenomenon; however, variation in the venom of peptidome is poorly understood [9,10]. Knowledge of variation of peptidome variation is of prime importance as these are the most potent toxins and less immunogenic [10,11]. Recently, it was shown that the commercial available antivenoms are ineffective in neutralizing the toxic and lethal effects of the peptides fraction and the antibody raised against this fraction was found to cross-react with all the regional venoms [11]. Further, peptides are known to be more potent in toxic and lethal effects [9,12]. Therefore studies on the variation in the venom of peptidome are of prime importance for development of efficacious antivenom. Several approaches and techniques have been employed for studying the variability of venom components that influences the pharmacological effects. Mass spectrometry is one of the major investigative tools, known to gives access to wealth of information in a short working time frame and with minute amount of samples as in the case of snake venoms [13]. Several studies have used MALDITOF to understand the influence of variation in snake venom induced pathophysiological effects [10,13-16].Indian cobra (Naja naja), one of the medically important snakes, is endemic and distributed all across the country. It is responsible for large number of morbidity and mortality cases in India [17]. It is observed that the variation in the venom composition of Naja naja species was the main reason for severity of pathogenesis in the victims of three districts of West Bengal (Eastern India) and the available polyvalent antivenom manufactured in western India was hardly effective in neutralizing the pathobiological manifestation of the venom [7]. The venom of Naja naja species from different geographical regions are known to vary in their biochemical and pharmacological activities and eastern regional venom is known to be the most toxic and lethal among the three regional venoms [18,19]. It was concluded that the variability in toxicity and lethality was due to the presence of peptides and phospholipases A2 (low molecular weight factors) in the venom [7,19]. Although intra-specific variability in Naja naja species venom has been observed [7,8,18,19], its influence on antivenom production is poorly understood, due to lack of characterization of major toxins involved in pharmacological effects. It is believed that an antiserum raised against fractionated venom containing such major toxins, could yield better protection [20]. Therefore, the aim of the study is to obtain general information of the peptidome (Low Molecular Weight fraction) - main toxic/ lethal component of the venom of Naja naja species from different regions of India, which would help in as a reference for development of efficacious therapeutic antivenom. Here we report for the first time the comparative peptidomic analysis of different regional venoms using MALDI-TOF for its potential use in efficacious therapeutic antivenom.
Chemicals and Drugs
MaterialsAll the reagents used were of proteomic grade. The lyophilized venom from Naja naja from different regions of India was a gift from Dr. T. Veerabasappa Gowda (Professor, University of Mysore), matrices of MALDI mass spectrometry; α- cyano-4-hydroxycinnamic acid in case of peptides and sinapinic acid in case of proteins used were from sigma-Aldrich (St. Louis, USA). Mass spectrometry calibration standards were from sigma-Aldrich (St. Louis, USA).
Extraction of low molecular weight fraction form Naja naja venoms
To obtain peptidome fraction (low molecular weight fractions), all the three regional Naja naja venom samples were separately subjected to G-50 column chromatography which was equilibrated and eluted with 0.1 M phosphate buffer (pH 7.0) containing 0.5 M Nacl. The elution resulted in 2 peaks out of which the 1st peak contained high molecular weight proteins (>50 KDa) and 2nd peak which contained low molecular weight proteins The samples were lyophilized until Mass spectrometric analysis was performed.
MALDI mass spectrometry of low molecular weight fractions
MALDI spectra were recorded in positive ion mode using a Bruker Daltonics Ultraflex TOF/TOF spectrometer. The matrices used for positive ion mode detection were α-cyano-4-hydroxycinnamic acid in 60% acetonitrile containing 0.1% TFA. Routinely, 0.5 μl of matrix was mixed with 0.5 μl (1 μg dissolved in 10 μl of water) of the peptide sample on a MALDI plate for mass spectral analysis. Each sample was spotted twice and spectra were recorded for each spot.
Results and Discussions
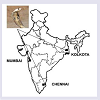
The venom of Indian Cobra (Naja naja) obtained from 3 different regions western (Mumbai, Maharastra), southern (Chennai, TamilNadu) and eastern (Kolkata, West Bengal) (Figure 1) varied greatly in MALDI-TOF spectral profile of peptide (low molecular weight) fractions in the 1-40 KDa window (Figure 2). The observed variations were in the presence/absence of different molecular weight peptides and their abundance across different regional venoms (Figure 2). The peptide pattern of the three regional venoms had similarities in containing 27 KDa, 20 KDa, 13 KDa and 6.7 KDa toxin components (Figure 2). In Elapidae venoms it is usually observed that 12-14 KDa members are PLA2s and < 7 KDa members is generally represented by neurotoxins, cardiotoxins or ion-channel blockers [20]. Further, it is known that ~5.7-7.3 KDa members are potential three finger toxins (3FTxs) [20]. Proteins from 3FTx and PLA2 families are generally known for the major biological effects, being responsible mainly for neurotoxicity and death by respiratory arrest which is the predominant clinical manifestation of Naja naja venoms and in general elapidae venoms [21-23].Although many isoforms of PLA2s (in the range of 12-14 KDa) have been isolated and characterized from these three regional venoms of Naja naja species, only few have been characterized [24-27]. A highly lethal cytotoxic peptide (~6.9 KDa) has been reported from eastern regional venom [28]. Recently, a potent cardiotoxin having Mol wt of 6.7 KDa has been reported [29].
The eastern and western regional venoms contained 12 KDa, 24 KDa and 33 KDa peptides which was completely absent in southern regional sample of Naja naja. However, it is interesting to note that the eastern regional venom sample contained an abundant ~5.7 KDa peptide which was completely absent in other regional venoms. This component might be the cardiotoxins or neurotoxins which predominantly belong to 3FTxs family that are least studied in Indian venoms. Supporting this is the observation that eastern region venom predominantly causes damage to cardiac muscle [8]. Further the study shows that a distinct PLA2 enzyme is known to be present in eastern venom, which is absent in southern and western venom samples [8]. Therefore the presence of highly potent low molecular weight components like the isoforms of PLA2s and 3FTxs might be the reason for the observed highly toxic and lethal effects of eastern regional venom when compared to other regional venoms [7,18,19]. Further, it can be viewed that the observed ineffectiveness of commercial polyvalent antivenom (manufactured in western India) in neutralizing the pathobiological manifestation of the venom samples from eastern India, might be due to its ineffectiveness of action on low molecular weight components’ like 3FTs and neurotoxins [7].
The absence of 12 KDa, 24 KDa and 33 KDa peptides in southern venom when compared to other regional venoms and the abundant existence of ~5.7 KDa peptide in only eastern regional venom of Naja naja species could well be utilized for selection of specimens in the production of antivenom and thus used in the treatment of snakebite patents. A similar study demonstrated and discussed the implications of mass spectrometry analysis for the production of Micrurus antivenoms [20]. Further, it concluded that an antiserum raised against fractionated venom containing such major toxins, could yield better protection [20].
Conclusion
In conclusion, the MALDI-TOF analysis revealed qualitative and quantitative peptidome variation in Naja naja venom of distinct geographical origin. Further this study contributes to towards the understanding of intra-venom variability which is useful for production of efficacious and region-specific therapeutic antivenoms, which is the immediate medical concern for researches.Acknowledgements
The authors thank Prof. P. Balram and Prof. T. Verrabassapa Gowda for their support. DBL thank Jain University for the constant encouragement given to progress in research. Dhananjaya BL (DBL) thanks DST IndoSrilankan - DST- MTR (DST/INT/SLP/P-007/2012, dated 5th May, 2014), Indo-European - Marie Curie IRSES Grant - (PIRSES-GA-2013-612131, dated 15th Nov, 2014).References
- Aird SD (2002) Ophidian envenomation strategies and the role of purines. Toxicon 40: 335-393.
- Dhananjaya BL, D'Souza CJ (2010) The pharmacological role of nucleotidases in snake venoms. Cell Biochem Funct 28: 171-177.
- Chippaux JP, Williams V, White J (1991) Snake venom variability: methods of study, results and interpretation. Toxicon 29: 1279-1303.
- Calvete JJ, Juárez P, Sanz L (2007) Snake venomics. Strategy and applications. J Mass Spectrom 42: 1405-1414.
- Theakston RD, Warrell DA, Griffiths E (2003) Report of a WHO workshop on the standardization and control of antivenoms. Toxicon 41: 541-557.
- Phillips RE, Theakston RD, Warrell DA, Galigedara Y, Abeysekera DT, et al. (1988) Paralysis, rhabdomyolysis and haemolysis caused by bites of Russell`s viper (Vipera russelli pulchella) in Sri Lanka: failure of Indian (Haffkine) antivenom. Q J Med 68: 691-715.
- Mukherjee AK, Maity CR (1998) The composition of Naja naja venom samples from three districts of West Bengal, India. Comp Biochem Physiol A Mol Integr Physiol 119: 621-627.
- Shashidharamurthy R, Mahadeswaraswamy YH, Ragupathi L, Vishwanath BS, Kemparaju K (2010) Systemic pathological effects induced by cobra (Naja naja) venom from geographically distinct origins of Indian peninsula. Exp Toxicol Pathol 62: 587-592.
- Rates B, Ferraz KK, Borges MH, Richardson M, De Lima ME (2008) Tityus serrulatus venom peptidomics: assessing venom peptide diversity. Toxicon 52: 611-618.
- Zelanis A, Tashima AK, Rocha MM, Furtado MF, Camargo AC (2010) Analysis of the ontogenetic variation in the venom proteome/peptidome of Bothrops jararaca reveals different strategies to deal with prey. J Proteome Res 9: 2278-2291.
- Kumar AV, Gowda TV (2006) Novel non-enzymatic toxic peptide of Daboia russelii (Eastern region) venom renders commercial polyvalent antivenom ineffective. Toxicon 47: 398-408.
- Calvete JJ, Sanz L, Angulo Y, Lomonte B, Gutiérrez JM (2009) Venoms, venomics, antivenomics. FEBS Lett 583: 1736-1743.
- Fox JW, Serrano SM (2008) Exploring snake venom proteomes: multifaceted analysis for complex toxin mixtures. Proteomics 8: 909-920.
- Creer S, Malhotra A, Thorpe RS, Stöcklin RS, Favreau PS, et al. (2003) Genetic and ecological correlates of intraspecific variation in pitviper venom composition detected using matrix-assisted laser desorption time-of-flight mass spectrometry (MALDI-TOF-MS) and isoelectric focusing. J Mol Evol 56: 317-329.
- Yanes O, Avilés FX, Wenzel R, Nazabal A, Zenobi R, et al. (2007) Proteomic profiling of a snake venom using high mass detection MALDI-TOF mass spectrometry. J Am Soc Mass Spectrom 18: 600-606.
- Escoubas P, Quinton L, Nicholson GM (2008) Venomics: unravelling the complexity of animal venoms with mass spectrometry. J Mass Spectrom 43: 279-295.
- Alirol E, Sharma SK, Bawaskar HS, Kuch U, Chappuis F (2010) Snake bite in South Asia: a review. PLoS Negl Trop Dis 4: e603.
- Shashidharamurthy R, Jagadeesha DK, Girish KS, Kemparaju K (2002) Variations in biochemical and pharmacological properties of Indian cobra (Naja naja naja) venom due to geographical distribution. Mol Cell Biochem 229: 93-101.
- Shashidharamurthy R, Kemparaju K (2007) Region-specific neutralization of Indian cobra (Naja naja) venom by polyclonal antibody raised against the eastern regional venom: a comparative study of the venoms from the three different geographical distributions. Int Immunopharmacol 7: 61-69.
- Ciscotto PH, Rates B, Silva DA, Richardson M, Silva LP, et al. (2011) Venomic analysis and evaluation of antivenom cross-reactivity of South American Micrurus species. J Proteomics 74: 1810-1825.
- Kini RM (1997) Phospholipase A2: a complex multifunctional protein puzzle. In: Kini RM (ed). Venom Phospholipase A2 Enzymes: structure, function and mechanism. Chichester, England: John Wiley pp.1-28.
- Doley R, Zhou X, Kini RM (2009) Snake venom phospholipase A2 enzymes. In: Mackessy SP (ed). Handbook of venoms and toxins of reptiles. CRC Press, Boca Raton, Florida pp.173-215.
- Doley R, Hegde RP, Rajagopalan N, Doley R, Kini RM (2009) Snake venom three-finger toxins. In: Mackessy SP (ed). Handbook of venoms and toxins of reptiles. CRC Press, Boca Raton. Florida pp.287-303.
- Doley R, Hegde Bhat MK, Gowda TV (1991) Isolation and characterization of a lethal phospholipase A2 (NN-IVb1-PLA2) from the Indian cobra (Naja naja naja) venom. Biochem Int 25: 1023-1034.
- Satish S, Tejaswini J, Krishnakantha TP, Gowda TV (2004) Purification of a class B1 platelet aggregation inhibitor phospholipase A2 from Indian cobra (Naja naja) venom. Biochimie 86: 203-210.
- Machiah DK, Gowda TV (2006) Purification of a post-synapic neurotoxic phospholipase A2 from Naja naja venom and its inhibition by a glycoprotein from Withania somnifera. Biochimie 88: 701-710.
- Shashidharamurthy R, Kemparaju K (2006) A neurotoxic phospholipase A2 variant; isolation and characterization from eastern regional Indian cobra (Naja naja) venom. Toxicon 47: 727-733.
- Ponnappa KC, Saviour P, Ramachandra NB, Kini RM, Gowda TV (2008) INN-toxin, a highly lethal peptide from the venom of Indian cobra (Naja naja) venom-isolation, characterization and pharmacological actions. Peptides 29: 1893-1900.
- Das T, Bhattacharya S, Halder B, Biswas A, Das Gupta S, et al. (2011) Cytotoxic and antioxidant property of a purified fraction (NN-32) of Indian Naja naja venom on Ehrlich ascites carcinoma in BALB/c mice. Toxicon 57: 1065-1072.